The (52-96) C-terminal domain of Vpr stimulates HIV-1 IN-mediated homologous strand transfer of mini-viral DNA
- PMID: 12736319
- PMCID: PMC156046
- DOI: 10.1093/nar/gkg364
The (52-96) C-terminal domain of Vpr stimulates HIV-1 IN-mediated homologous strand transfer of mini-viral DNA
Abstract
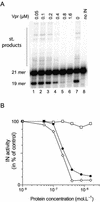
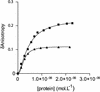
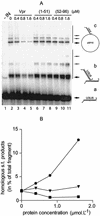
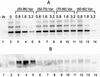
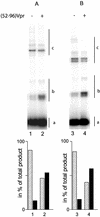
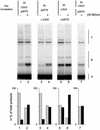
Viral integrase (IN) and Vpr are both components of the human immunodeficiency virus type 1 (HIV-1) pre-integration complex. To investigate whether these proteins interact within this complex, we investigated the effects of Vpr and its subdomains on IN activity in vitro. When a 21mer oligonucleotide was used as a donor and acceptor, both Vpr and its C-terminal DNA-binding domain [(52-96)Vpr] inhibited the integration reaction, whereas the (1-51)Vpr domain did not affect IN activity. Steady-state fluorescence anisotropy showed that both full-length and (52-96)Vpr bind to the short oligonucleotide, thereby extending previous observations with long DNA. The concentrations of the two proteins required to inhibit IN activity were consistent with their affinities for the oligonucleotide. The use of a 492 bp mini-viral substrate confirmed that Vpr can inhibit the IN-mediated reaction. However, the activity of (52-96)Vpr differed notably since it stimulated specifically integration events involving two homologous mini-viral DNAs. Order of addition experiments indicated that the stimulation was maximal when IN, (50-96)Vpr and the mini-viral DNA were allowed to form a complex. Furthermore, in the presence of (50-96)Vpr, the binding of IN to the mini-viral DNA was dramatically enhanced. Taken together, these data suggest that (52-96)Vpr stimulates the formation of a specific complex between IN and the mini-viral DNA.
Figures

Similar articles
-
HIV-1 Vpr binding to HIV-1 LTR C/EBP cis-acting elements and adjacent regions is sequence-specific.Biomed Pharmacother. 2003 Jan;57(1):41-8. doi: 10.1016/s0753-3322(02)00333-5. Biomed Pharmacother. 2003. PMID: 12642036
-
A novel Vpr peptide interactor fused to integrase (IN) restores integration activity to IN-defective HIV-1 virions.Virology. 1999 Mar 1;255(1):77-85. doi: 10.1006/viro.1998.9544. Virology. 1999. PMID: 10049823
-
Inhibition of the activities of reverse transcriptase and integrase of human immunodeficiency virus type-1 by peptides derived from the homologous viral protein R (Vpr).J Mol Biol. 2007 Jun 22;369(5):1230-43. doi: 10.1016/j.jmb.2007.03.073. Epub 2007 Apr 1. J Mol Biol. 2007. PMID: 17490682
-
Partner molecules of accessory protein Vpr of the human immunodeficiency virus type 1.DNA Cell Biol. 2004 Apr;23(4):193-205. doi: 10.1089/104454904773819789. DNA Cell Biol. 2004. PMID: 15142377 Review.
-
HIV-1 Vpr: genetic diversity and functional features from the perspective of structure.DNA Cell Biol. 2004 Apr;23(4):207-22. doi: 10.1089/104454904773819798. DNA Cell Biol. 2004. PMID: 15142378 Review.
Cited by
-
Cell-permeable stapled peptides based on HIV-1 integrase inhibitors derived from HIV-1 gene products.ACS Chem Biol. 2013 Oct 18;8(10):2235-44. doi: 10.1021/cb400495h. Epub 2013 Aug 15. ACS Chem Biol. 2013. PMID: 23898787 Free PMC article.
-
HIV-1 IN inhibitors: 2010 update and perspectives.Curr Top Med Chem. 2009;9(11):1016-37. doi: 10.2174/156802609789630910. Curr Top Med Chem. 2009. PMID: 19747122 Free PMC article. Review.
-
A protein ballet around the viral genome orchestrated by HIV-1 reverse transcriptase leads to an architectural switch: from nucleocapsid-condensed RNA to Vpr-bridged DNA.Virus Res. 2013 Feb;171(2):287-303. doi: 10.1016/j.virusres.2012.09.008. Epub 2012 Sep 24. Virus Res. 2013. PMID: 23017337 Free PMC article.
-
In vitro initial attachment of HIV-1 integrase to viral ends: control of the DNA specific interaction by the oligomerization state.Nucleic Acids Res. 2008 Dec;36(22):7043-58. doi: 10.1093/nar/gkn796. Epub 2008 Nov 5. Nucleic Acids Res. 2008. PMID: 18987001 Free PMC article.
-
The host-pathogen interaction of human cyclophilin A and HIV-1 Vpr requires specific N-terminal and novel C-terminal domains.BMC Struct Biol. 2011 Dec 20;11:49. doi: 10.1186/1472-6807-11-49. BMC Struct Biol. 2011. PMID: 22185200 Free PMC article.
References
-
- Fujiwara T. and Mizuuchi,K. (1988) Retroviral DNA integration: structure of an integration intermediate. Cell, 54, 497–504. - PubMed

