Substructure of direction-selective receptive fields in macaque V1
- PMID: 12740412
- PMCID: PMC2627773
- DOI: 10.1152/jn.00822.2002
Substructure of direction-selective receptive fields in macaque V1
Abstract
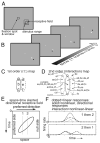
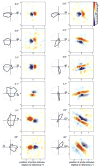
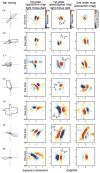
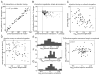
We used two-dimensional (2-D) sparse noise to map simultaneous and sequential two-spot interactions in simple and complex direction-selective cells in macaque V1. Sequential-interaction maps for both simple and complex cells showed preferred-direction facilitation and null-direction suppression for same-contrast stimulus sequences and the reverse for inverting-contrast sequences, although the magnitudes of the interactions were weaker for the simple cells. Contrast-sign selectivity in complex cells indicates that direction-selective interactions in these cells must occur in antecedent simple cells or in simple-cell-like dendritic compartments. Our maps suggest that direction selectivity, and on and off segregation perpendicular to the orientation axis, can occur prior to receptive-field elongation along the orientation axis. 2-D interaction maps for some complex cells showed elongated alternating facilitatory and suppressive interactions as predicted if their inputs were orientation-selective simple cells. The negative interactions, however, were less elongated than the positive interactions, and there was an inflection at the origin in the positive interactions, so the interactions were chevron-shaped rather than band-like. Other complex cells showed only two round interaction regions, one negative and one positive. Several explanations for the map shapes are considered, including the possibility that directional interactions are generated directly from unoriented inputs.
Figures





Similar articles
-
Selectivity and spatial distribution of signals from the receptive field surround in macaque V1 neurons.J Neurophysiol. 2002 Nov;88(5):2547-56. doi: 10.1152/jn.00693.2001. J Neurophysiol. 2002. PMID: 12424293
-
Receptive fields of disparity-selective neurons in macaque striate cortex.Nat Neurosci. 1999 Sep;2(9):825-32. doi: 10.1038/12199. Nat Neurosci. 1999. PMID: 10461222
-
Receptive fields and functional architecture of macaque V2.J Neurophysiol. 1994 Jun;71(6):2517-42. doi: 10.1152/jn.1994.71.6.2517. J Neurophysiol. 1994. PMID: 7931532
-
Surround suppression supports second-order feature encoding by macaque V1 and V2 neurons.Vision Res. 2014 Nov;104:24-35. doi: 10.1016/j.visres.2014.10.004. Epub 2014 Oct 23. Vision Res. 2014. PMID: 25449336 Free PMC article. Review.
-
The contribution of vertical and horizontal connections to the receptive field center and surround in V1.Neural Netw. 2004 Jun-Jul;17(5-6):681-93. doi: 10.1016/j.neunet.2004.05.002. Neural Netw. 2004. PMID: 15288892 Review.
Cited by
-
Direct Measurement of Correlation Responses in Drosophila Elementary Motion Detectors Reveals Fast Timescale Tuning.Neuron. 2016 Oct 5;92(1):227-239. doi: 10.1016/j.neuron.2016.09.017. Neuron. 2016. PMID: 27710784 Free PMC article.
-
Neural basis for a powerful static motion illusion.J Neurosci. 2005 Jun 8;25(23):5651-6. doi: 10.1523/JNEUROSCI.1084-05.2005. J Neurosci. 2005. PMID: 15944393 Free PMC article.
-
Mapping of visual receptive fields by tomographic reconstruction.Neural Comput. 2012 Oct;24(10):2543-78. doi: 10.1162/NECO_a_00334. Epub 2012 Jun 26. Neural Comput. 2012. PMID: 22734491 Free PMC article.
-
The Neuronal Basis of an Illusory Motion Percept Is Explained by Decorrelation of Parallel Motion Pathways.Curr Biol. 2018 Dec 3;28(23):3748-3762.e8. doi: 10.1016/j.cub.2018.10.007. Epub 2018 Nov 21. Curr Biol. 2018. PMID: 30471993 Free PMC article.
-
Computational diversity in complex cells of cat primary visual cortex.J Neurosci. 2007 Sep 5;27(36):9638-48. doi: 10.1523/JNEUROSCI.2119-07.2007. J Neurosci. 2007. PMID: 17804624 Free PMC article.
References
-
- Adelson EH, Bergen JR. Spatiotemporal energy models for the perception of motion. J Opt Soc Am A Opt and Image Sci. 1985;2:284–299. - PubMed
-
- Albright TD, Desimone R, Gross CG. Columnar organization of directionally selective cells in visual area MT of the macaque. J Neurophysiol. 1984;51:16–31. - PubMed
-
- Anstis SM, Rogers BJ. Illusory reversal of visual depth and movement during changes of contrast. Vis Res. 1975;15:957–961. - PubMed
-
- Anzai A, Ohzawa I, Freeman RD. Neural mechanisms for processing binocular information. II. Complex cells. J Neurophysiol. 1999;82:909–924. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

