Surfactant proteins A and D inhibit the growth of Gram-negative bacteria by increasing membrane permeability
- PMID: 12750409
- PMCID: PMC155045
- DOI: 10.1172/JCI16889
Surfactant proteins A and D inhibit the growth of Gram-negative bacteria by increasing membrane permeability
Abstract
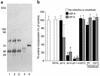
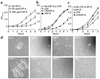
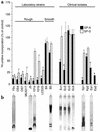
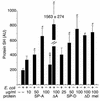
The pulmonary collectins, surfactant proteins A (SP-A) and D (SP-D), have been reported to bind lipopolysaccharide (LPS), opsonize microorganisms, and enhance the clearance of lung pathogens. In this study, we examined the effect of SP-A and SP-D on the growth and viability of Gram-negative bacteria. The pulmonary clearance of Escherichia coli K12 was reduced in SP-A-null mice and was increased in SP-D-overexpressing mice, compared with strain-matched wild-type controls. Purified SP-A and SP-D inhibited bacterial synthetic functions of several, but not all, strains of E. coli, Klebsiella pneumoniae, and Enterobacter aerogenes. In general, rough E. coli strains were more susceptible than smooth strains, and collectin-mediated growth inhibition was partially blocked by coincubation with rough LPS vesicles. Although both SP-A and SP-D agglutinated E. coli K12 in a calcium-dependent manner, microbial growth inhibition was independent of bacterial aggregation. At least part of the antimicrobial activity of SP-A and SP-D was localized to their C-terminal domains using truncated recombinant proteins. Incubation of E. coli K12 with SP-A or SP-D increased bacterial permeability. Deletion of the E. coli OmpA gene from a collectin-resistant smooth E. coli strain enhanced SP-A and SP-D-mediated growth inhibition. These data indicate that SP-A and SP-D are antimicrobial proteins that directly inhibit the proliferation of Gram-negative bacteria in a macrophage- and aggregation-independent manner by increasing the permeability of the microbial cell membrane.
Figures
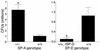
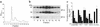
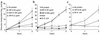




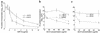

Comment in
-
Pulmonary surfactant: a front line of lung host defense.J Clin Invest. 2003 May;111(10):1453-5. doi: 10.1172/JCI18650. J Clin Invest. 2003. PMID: 12750392 Free PMC article. Review. No abstract available.
References
-
- Medzhitov R, Janeway CA., Jr Innate immunity: the virtues of a nonclonal system of recognition. Cell. 1997;91:295–298. - PubMed
-
- Hoffmann JA, Kafatos FC, Janeway CA, Ezekowitz RA. Phylogenetic perspectives in innate immunity. Science. 1999;284:1313–1318. - PubMed
-
- Wright JR. Immunomodulatory functions of surfactant. Physiol. Rev. 1997;77:931–962. - PubMed
-
- Drickamer K, Dordal MS, Reynolds L. Mannose-binding proteins isolated from rat liver contain carbohydrate-recognition domains linked to collagenous tails. Complete primary structures and homology with pulmonary surfactant apoprotein. J. Biol. Chem. 1986;261:6878–6887. - PubMed
-
- Crouch EC. Collectins and pulmonary host defense. Am. J. Respir. Cell Mol. Biol. 1998;19:177–201. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

