Reducing the computational complexity of protein folding via fragment folding and assembly
- PMID: 12761388
- PMCID: PMC2323902
- DOI: 10.1110/ps.0232903
Reducing the computational complexity of protein folding via fragment folding and assembly
Abstract
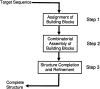
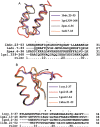
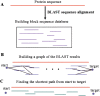
Understanding, and ultimately predicting, how a 1-D protein chain reaches its native 3-D fold has been one of the most challenging problems during the last few decades. Data increasingly indicate that protein folding is a hierarchical process. Hence, the question arises as to whether we can use the hierarchical concept to reduce the practically intractable computational times. For such a scheme to work, the first step is to cut the protein sequence into fragments that form local minima on the polypeptide chain. The conformations of such fragments in solution are likely to be similar to those when the fragments are embedded in the native fold, although alternate conformations may be favored during the mutual stabilization in the combinatorial assembly process. Two elements are needed for such cutting: (1) a library of (clustered) fragments derived from known protein structures and (2) an assignment algorithm that selects optimal combinations to "cover" the protein sequence. The next two steps in hierarchical folding schemes, not addressed here, are the combinatorial assembly of the fragments and finally, optimization of the obtained conformations. Here, we address the first step in a hierarchical protein-folding scheme. The input is a target protein sequence and a library of fragments created by clustering building blocks that were generated by cutting all protein structures. The output is a set of cutout fragments. We briefly outline a graph theoretic algorithm that automatically assigns building blocks to the target sequence, and we describe a sample of the results we have obtained.
Figures







Similar articles
-
Hierarchical protein folding pathways: a computational study of protein fragments.Proteins. 2003 May 1;51(2):203-15. doi: 10.1002/prot.10294. Proteins. 2003. PMID: 12660989
-
Comparison of protein fragments identified by limited proteolysis and by computational cutting of proteins.Protein Sci. 2002 Jul;11(7):1753-70. doi: 10.1110/ps.4100102. Protein Sci. 2002. PMID: 12070328 Free PMC article.
-
Folding stability and cooperativity of the three forms of 1-110 residues fragment of staphylococcal nuclease.Biophys J. 2007 Mar 15;92(6):2090-107. doi: 10.1529/biophysj.106.092155. Epub 2006 Dec 15. Biophys J. 2007. PMID: 17172296 Free PMC article.
-
Protein folding: binding of conformationally fluctuating building blocks via population selection.Crit Rev Biochem Mol Biol. 2001;36(5):399-433. doi: 10.1080/20014091074228. Crit Rev Biochem Mol Biol. 2001. PMID: 11724155 Review.
-
Knowledge-based potential functions in protein design.Curr Opin Struct Biol. 2002 Aug;12(4):447-52. doi: 10.1016/s0959-440x(02)00346-9. Curr Opin Struct Biol. 2002. PMID: 12163066 Review.
Cited by
-
Elucidating quantitative stability/flexibility relationships within thioredoxin and its fragments using a distance constraint model.J Mol Biol. 2006 May 5;358(3):882-904. doi: 10.1016/j.jmb.2006.02.015. Epub 2006 Feb 24. J Mol Biol. 2006. PMID: 16542678 Free PMC article.
-
Possibilities of Using De Novo Design for Generating Diverse Functional Food Enzymes.Int J Mol Sci. 2023 Feb 14;24(4):3827. doi: 10.3390/ijms24043827. Int J Mol Sci. 2023. PMID: 36835238 Free PMC article. Review.
-
Designing succinct structural alphabets.Bioinformatics. 2008 Jul 1;24(13):i182-9. doi: 10.1093/bioinformatics/btn165. Bioinformatics. 2008. PMID: 18586712 Free PMC article.
-
Protein fragments: functional and structural roles of their coevolution networks.PLoS One. 2012;7(11):e48124. doi: 10.1371/journal.pone.0048124. Epub 2012 Nov 5. PLoS One. 2012. PMID: 23139761 Free PMC article.
-
Probing protein fold space with a simplified model.J Mol Biol. 2008 Jan 25;375(4):920-33. doi: 10.1016/j.jmb.2007.10.087. Epub 2007 Nov 9. J Mol Biol. 2008. PMID: 18054792 Free PMC article.
References
-
- Abagyan, R.A. and Batalov, S. 1997. Do aligned sequences share the same fold? J. Mol. Biol. 273 355–368. - PubMed
-
- Altschul, S.F., Gish, W., Miller, W., Myers, E.W., and Lipman, D.J. 1990. Basic local alignment search tool. J. Mol. Biol. 215 403–410. - PubMed
-
- Baldwin, R.L. and Rose, G.D. 1999a. Is protein folding hierarchic? I. Local structure and peptide folding. Trends Biochem. Sci. 24 26–33. - PubMed
-
- ———. 1999b. Is protein folding hierarchic? II. Folding intermediates and transition states. Trends Biochem. Sci. 24 77–84. - PubMed
-
- Bernstein, F.C., Koetzle, T.F., Williams, G.J.B., Meyer, E.F., Brice, M.D., Rodgers, J.R., Kennard, O., Shimanouchi, T., and Tasumi, M. 1977. The Protein Data Bank: A computer-based archival file for macromolecular structures. J. Mol. Biol. 112 535–542. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

