Flavivirus capsid is a dimeric alpha-helical protein
- PMID: 12768036
- PMCID: PMC156156
- DOI: 10.1128/jvi.77.12.7143-7149.2003
Flavivirus capsid is a dimeric alpha-helical protein
Abstract
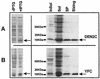
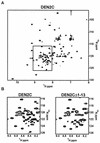
The capsid proteins of two flaviviruses, yellow fever virus and dengue virus, were expressed in Escherichia coli and purified to near homogeneity suitable for biochemical characterization and structure determination by nuclear magnetic resonance. The oligomeric properties of the capsid protein in solution were investigated. In the absence of nucleic acid, both proteins were predominantly dimeric in solution. Further analysis of both proteins with far-UV circular dichroism spectroscopy indicated that they were largely alpha-helical. The secondary structure elements of the dengue virus capsid were determined by chemical shift indexing of the sequence-specific backbone resonance assignments. The dengue virus capsid protein devoid of its C-terminal signal sequence was found to be composed of four alpha helices. The longest alpha helix, 20 residues, is located at the C terminus and has an amphipathic character. In contrast, the N terminus was found to be unstructured and could be removed without disrupting the structural integrity of the protein.
Figures

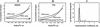

Similar articles
-
The 1H, 15N and 13C resonance assignments of dengue virus capsid protein with the deletion of the intrinsically disordered N-terminal region.Biomol NMR Assign. 2023 Jun;17(1):23-26. doi: 10.1007/s12104-022-10115-1. Epub 2023 Feb 1. Biomol NMR Assign. 2023. PMID: 36723824
-
Unlike dengue virus, the conserved 14-23 residues in N-terminal region of Zika virus capsid is not involved in lipid interactions.Biochim Biophys Acta Biomembr. 2020 Nov 1;1862(11):183440. doi: 10.1016/j.bbamem.2020.183440. Epub 2020 Aug 9. Biochim Biophys Acta Biomembr. 2020. PMID: 32783888
-
Structural and Functional Properties of the Capsid Protein of Dengue and Related Flavivirus.Int J Mol Sci. 2019 Aug 8;20(16):3870. doi: 10.3390/ijms20163870. Int J Mol Sci. 2019. PMID: 31398956 Free PMC article.
-
Unique structural features of flaviviruses' capsid proteins: new insights on structure-function relationship.Curr Opin Virol. 2021 Apr;47:106-112. doi: 10.1016/j.coviro.2021.02.005. Epub 2021 Mar 12. Curr Opin Virol. 2021. PMID: 33721656 Review.
-
Structure-guided paradigm shifts in flavivirus assembly and maturation mechanisms.Adv Virus Res. 2020;108:33-83. doi: 10.1016/bs.aivir.2020.08.003. Epub 2020 Sep 23. Adv Virus Res. 2020. PMID: 33837721 Free PMC article. Review.
Cited by
-
Searching for drug leads targeted to the hydrophobic cleft of dengue virus capsid protein.J Enzyme Inhib Med Chem. 2022 Dec;37(1):287-298. doi: 10.1080/14756366.2021.2004591. J Enzyme Inhib Med Chem. 2022. PMID: 34894959 Free PMC article.
-
Structural insight into the Zika virus capsid encapsulating the viral genome.Cell Res. 2018 Apr;28(4):497-499. doi: 10.1038/s41422-018-0007-9. Epub 2018 Feb 21. Cell Res. 2018. PMID: 29467384 Free PMC article. No abstract available.
-
Characterization of the mode of action of a potent dengue virus capsid inhibitor.J Virol. 2014 Oct;88(19):11540-55. doi: 10.1128/JVI.01745-14. Epub 2014 Jul 23. J Virol. 2014. PMID: 25056895 Free PMC article.
-
A novel system for visualizing alphavirus assembly.J Virol Methods. 2015 Sep 15;222:158-63. doi: 10.1016/j.jviromet.2015.06.013. Epub 2015 Jun 27. J Virol Methods. 2015. PMID: 26122073 Free PMC article.
-
Capsid-Targeted Viral Inactivation: A Novel Tactic for Inhibiting Replication in Viral Infections.Viruses. 2016 Sep 21;8(9):258. doi: 10.3390/v8090258. Viruses. 2016. PMID: 27657114 Free PMC article. Review.
References
-
- Cantor, C. R., and P. R. Schimmel. 1980. Techniques for the study of biological structure and function. Part II. W. H. Freeman and Co., San Francisco, Calif.
-
- Delaglio, F., S. Grzesiek, G. W. Vuister, G. Zhu, J. Pfeifer, and A. Bax. 1995. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6:277-293. - PubMed
-
- Fonseca, B. A., S. Pincus, R. E. Shope, E. Paoletti, and P. W. Mason. 1994. Recombinant vaccinia viruses co-expressing dengue-1 glycoproteins prM and E induce neutralizing antibodies in mice. Vaccine 12:279-285. - PubMed
-
- Goddard, T. D., and D. G. Kneller. 1993. SPARKY3. University of California, San Francisco.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

