Protein structure change studied by hydrogen-deuterium exchange, functional labeling, and mass spectrometry
- PMID: 12773622
- PMCID: PMC165829
- DOI: 10.1073/pnas.1232301100
Protein structure change studied by hydrogen-deuterium exchange, functional labeling, and mass spectrometry
Abstract
An automated high-throughput, high-resolution deuterium exchange HPLC-MS method (DXMS) was used to extend previous hydrogen exchange studies on the position and energetic role of regulatory structure changes in hemoglobin. The results match earlier highly accurate but much more limited tritium exchange results, extend the analysis to the entire sequence of both hemoglobin subunits, and identify some energetically important changes. Allosterically sensitive amide hydrogens located at near amino acid resolution help to confirm the reality of local unfolding reactions and their use to evaluate resolved structure changes in terms of allosteric free energy.
Figures


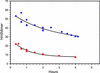

References
-
- Woodward, C. K. (1994) Curr. Opin. Struct. Biol. 4, 112-116.
-
- Scholtz, J. M. & Robertson, A. D. (1995) Methods Mol. Biol. 40, 291-311. - PubMed
-
- Raschke, T. M. & Marqusee, S. (1998) Curr. Opin. Biotechnol. 9, 80-86. - PubMed
-
- Rogero, J. R., Englander, J. J. & Englander, S. W. (1986) Methods Enzymol. 131, 508-517. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

