Developmentally regulated usage of Physarum DNA replication origins
- PMID: 12776736
- PMCID: PMC1319177
- DOI: 10.1038/sj.embor.embor822
Developmentally regulated usage of Physarum DNA replication origins
Abstract
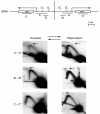
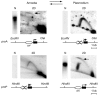
To determine the extent to which eukaryotic replication origins are developmentally regulated in transcriptionally competent cells, we compared origin use in untreated growing amoebae and plasmodia of Physarum polycephalum. At loci that contain genes transcribed in both developmental stages, such as the ribosomal RNA genes and two unlinked actin genes, we show that there is a similar replicational organization, with the same origins used with comparable efficiencies, as shown by two-dimensional agarose-gel electrophoresis. By contrast, we found cell-type-specific replication patterns for the homologous, unlinked profilin A (proA) and profilin P (proP) genes. proA is replicated from a promoter-proximal origin in amoebae, in which it is highly expressed, and is replicated passively in the plasmodium, in which it is not expressed. Conversely, proP is replicated passively and is not expressed in amoebae, but coincides with an efficient origin when highly expressed in the plasmodium. Our results show a reprogramming of S phase that is linked to the reprogramming of transcription during Physarum cell differentiation. This is achieved by the use of two classes of promoter-associated replication origins: those that are constitutively active and those that are developmentally regulated. This suggests that replication origins, like genes, are under epigenetic control associated with cellular differentiation.
Figures


Similar articles
-
Mapping of a Physarum chromosomal origin of replication tightly linked to a developmentally-regulated profilin gene.Nucleic Acids Res. 1992 Jul 11;20(13):3309-15. doi: 10.1093/nar/20.13.3309. Nucleic Acids Res. 1992. PMID: 1630902 Free PMC article.
-
Two early replicated, developmentally controlled genes of Physarum display different patterns of DNA replication by two-dimensional agarose gel electrophoresis.Chromosoma. 1993 Sep;102(8):563-74. doi: 10.1007/BF00368349. Chromosoma. 1993. PMID: 7902230
-
Replicational organization of three weakly expressed loci in Physarum polycephalum.Nucleic Acids Res. 2002 Jun 1;30(11):2261-9. doi: 10.1093/nar/30.11.2261. Nucleic Acids Res. 2002. PMID: 12034812 Free PMC article.
-
Plasmodium development in the myxomycete Physarum polycephalum: genetic control and cellular events.Microbiology (Reading). 1995 Oct;141 ( Pt 10):2355-65. doi: 10.1099/13500872-141-10-2355. Microbiology (Reading). 1995. PMID: 7581996 Review. No abstract available.
-
Genetic Diversity in the mtDNA of Physarum polycephalum.Genes (Basel). 2023 Mar 2;14(3):628. doi: 10.3390/genes14030628. Genes (Basel). 2023. PMID: 36980901 Free PMC article. Review.
Cited by
-
Linker histone phosphorylation regulates global timing of replication origin firing.J Biol Chem. 2009 Jan 30;284(5):2823-2829. doi: 10.1074/jbc.M805617200. Epub 2008 Nov 17. J Biol Chem. 2009. PMID: 19015270 Free PMC article.
-
Replication-independent core histone dynamics at transcriptionally active loci in vivo.Genes Dev. 2005 Mar 15;19(6):677-82. doi: 10.1101/gad.1265205. Genes Dev. 2005. PMID: 15769942 Free PMC article.
-
Genomic specification and epigenetic regulation of eukaryotic DNA replication origins.EMBO J. 2004 Nov 10;23(22):4365-70. doi: 10.1038/sj.emboj.7600450. Epub 2004 Oct 28. EMBO J. 2004. PMID: 15510221 Free PMC article. Review.
-
RNase-dependent discontinuities associated with the crossovers of spontaneously formed joint DNA molecules in Physarum polycephalum.Chromosoma. 2010 Dec;119(6):601-11. doi: 10.1007/s00412-010-0281-x. Epub 2010 Jul 7. Chromosoma. 2010. PMID: 20607271
-
Eukaryotic DNA replication origins: many choices for appropriate answers.Nat Rev Mol Cell Biol. 2010 Oct;11(10):728-38. doi: 10.1038/nrm2976. Nat Rev Mol Cell Biol. 2010. PMID: 20861881 Review.
References
-
- Bénard M., Maric C. & Pierron G. (2001) DNA replication-dependent formation of joint DNA molecules in Physarum polycephalum. Mol. Cell, 7, 971–980. - PubMed
-
- Binette F., Bénard M., Laroche A., Pierron G., Lemieux G. & Pallotta D. (1990) Cellspecific expression of a profilin gene family. DNA Cell Biol., 9, 323–334. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

