Localized recruitment of a chromatin-remodeling activity by an activator in vivo drives transcriptional elongation
- PMID: 12782657
- PMCID: PMC196071
- DOI: 10.1101/gad.1071803
Localized recruitment of a chromatin-remodeling activity by an activator in vivo drives transcriptional elongation
Abstract
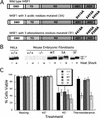
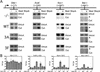
To understand the role of chromatin-remodeling activities in transcription, it is necessary to understand how they interact with transcriptional activators in vivo to regulate the different steps of transcription. Human heat shock factor 1 (HSF1) stimulates both transcriptional initiation and elongation. We replaced mouse HSF1 in fibroblasts with wild-type and mutant human HSF1 constructs and characterized regulation of an endogenous mouse hsp70 gene. A mutation that diminished transcriptional initiation led to twofold reductions in hsp70 mRNA induction and recruitment of a SWI/SNF remodeling complex. In contrast, a mutation that diminished transcriptional elongation abolished induction of full-length mRNA, SWI/SNF recruitment, and chromatin remodeling, but minimally impaired initiation from the hsp70 promoter. Another remodeling factor, SNF2h, is constitutively present at the promoter irrespective of the genotype of HSF1. These data suggest that localized recruitment of SWI/SNF drives a specialized remodeling reaction necessary for the production of full-length hsp70 mRNA.
Figures
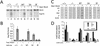


Similar articles
-
Heat shock transcription factor (Hsf)-4b recruits Brg1 during the G1 phase of the cell cycle and regulates the expression of heat shock proteins.J Cell Biochem. 2006 Aug 15;98(6):1528-42. doi: 10.1002/jcb.20865. J Cell Biochem. 2006. PMID: 16552721
-
[SWI/SNF Protein Complexes Participate in the Initiation and Elongation Stages of Drosophila hsp70 Gene Transcription].Genetika. 2016 Feb;52(2):164-9. Genetika. 2016. PMID: 27215030 Russian.
-
Heat shock factor-1 protein in heat shock factor-1 gene-transfected human epidermoid A431 cells requires phosphorylation before inducing heat shock protein-70 production.J Clin Invest. 1997 Jan 1;99(1):136-43. doi: 10.1172/JCI119124. J Clin Invest. 1997. PMID: 9011567 Free PMC article.
-
Interplay between chromatin modifying and remodeling complexes in transcriptional regulation.Crit Rev Eukaryot Gene Expr. 1999;9(3-4):221-30. doi: 10.1615/critreveukargeneexpr.v9.i3-4.70. Crit Rev Eukaryot Gene Expr. 1999. PMID: 10651239 Review.
-
Cell-free systems to study chromatin remodeling.Methods Cell Biol. 1998;53:497-515. doi: 10.1016/s0091-679x(08)60892-4. Methods Cell Biol. 1998. PMID: 9348522 Review. No abstract available.
Cited by
-
The SWI/SNF chromatin remodeling complex influences transcription by RNA polymerase I in Saccharomyces cerevisiae.PLoS One. 2013;8(2):e56793. doi: 10.1371/journal.pone.0056793. Epub 2013 Feb 20. PLoS One. 2013. PMID: 23437238 Free PMC article.
-
IKAROS is required for the measured response of NOTCH target genes upon external NOTCH signaling.PLoS Genet. 2021 Mar 26;17(3):e1009478. doi: 10.1371/journal.pgen.1009478. eCollection 2021 Mar. PLoS Genet. 2021. PMID: 33770102 Free PMC article.
-
Control of V(D)J Recombination through Transcriptional Elongation and Changes in Locus Chromatin Structure and Nuclear Organization.Genet Res Int. 2011;2011:970968. doi: 10.4061/2011/970968. Epub 2011 Sep 29. Genet Res Int. 2011. PMID: 22567371 Free PMC article.
-
T helper type 1-specific Brg1 recruitment and remodeling of nucleosomes positioned at the IFN-gamma promoter are Stat4 dependent.J Exp Med. 2006 Jun 12;203(6):1493-505. doi: 10.1084/jem.20060066. Epub 2006 May 22. J Exp Med. 2006. PMID: 16717115 Free PMC article.
-
The Swi/Snf complex is important for histone eviction during transcriptional activation and RNA polymerase II elongation in vivo.Mol Cell Biol. 2007 Oct;27(20):6987-95. doi: 10.1128/MCB.00717-07. Epub 2007 Aug 20. Mol Cell Biol. 2007. PMID: 17709398 Free PMC article.
References
-
- Aalfs J.D., Narlikar, G.J., and Kingston, R.E. 2001. Functional differences between the human ATP-dependent nucleosome remodeling proteins BRG1 and SNF2H. J. Biol. Chem. 276: 34270–34278. - PubMed
-
- Agalioti T., Lomvardas, S., Parekh, B., Yie, J., Maniatis, T., and Thanos, D. 2000. Ordered recruitment of chromatin modifying and general transcription factors to the IFN-β promoter. Cell 103: 667–678. - PubMed
-
- Ausubel F.M., Brent, R., Kingston, R.E., Moore, D.D., Seidman, J.G., Smith, J.A., and Struhl, K. 1996. Current protocols in molecular biology. John Wiley and Sons, New York.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
