NMR structure of the active conformation of the Varkud satellite ribozyme cleavage site
- PMID: 12782785
- PMCID: PMC165820
- DOI: 10.1073/pnas.0832440100
NMR structure of the active conformation of the Varkud satellite ribozyme cleavage site
Abstract
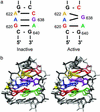
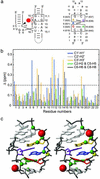
Substrate cleavage by the Neurospora Varkud satellite (VS) ribozyme involves a structural change in the stem-loop I substrate from an inactive to an active conformation. We have determined the NMR solution structure of a mutant stem-loop I that mimics the active conformation of the cleavage site internal loop. This structure shares many similarities, but also significant differences, with the previously determined structures of the inactive internal loop. The active internal loop displays different base-pairing interactions and forms a novel RNA fold composed exclusively of sheared G-A base pairs. From chemical-shift mapping we identified two Mg2+ binding sites in the active internal loop. One of the Mg2+ binding sites forms in the active but not the inactive conformation of the internal loop and is likely important for catalysis. Using the structure comparison program mc-search, we identified the active internal loop fold in other RNA structures. In Thermus thermophilus 16S rRNA, this RNA fold is directly involved in a long-range tertiary interaction. An analogous tertiary interaction may form between the active internal loop of the substrate and the catalytic domain of the VS ribozyme. The combination of NMR and bioinformatic approaches presented here has identified a novel RNA fold and provides insights into the structural basis of catalytic function in the Neurospora VS ribozyme.
Figures
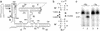
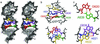
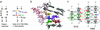
Similar articles
-
Nuclear magnetic resonance structure of the Varkud satellite ribozyme stem-loop V RNA and magnesium-ion binding from chemical-shift mapping.Biochemistry. 2005 Mar 22;44(11):4157-70. doi: 10.1021/bi047963l. Biochemistry. 2005. PMID: 15766243
-
A remarkably stable kissing-loop interaction defines substrate recognition by the Neurospora Varkud Satellite ribozyme.RNA. 2014 Sep;20(9):1451-64. doi: 10.1261/rna.046144.114. Epub 2014 Jul 22. RNA. 2014. PMID: 25051972 Free PMC article.
-
The NMR structure of the II-III-VI three-way junction from the Neurospora VS ribozyme reveals a critical tertiary interaction and provides new insights into the global ribozyme structure.RNA. 2015 Sep;21(9):1621-32. doi: 10.1261/rna.052076.115. Epub 2015 Jun 29. RNA. 2015. PMID: 26124200 Free PMC article.
-
The Varkud satellite ribozyme.RNA. 2004 Feb;10(2):151-8. doi: 10.1261/rna.5217104. RNA. 2004. PMID: 14730013 Free PMC article. Review.
-
The Neurospora Varkud satellite ribozyme.Biochem Soc Trans. 2002 Nov;30(Pt 6):1122-6. doi: 10.1042/bst0301122. Biochem Soc Trans. 2002. PMID: 12440987 Review.
Cited by
-
DNA mimicry by a high-affinity anti-NF-kappaB RNA aptamer.Nucleic Acids Res. 2008 Mar;36(4):1227-36. doi: 10.1093/nar/gkm1141. Epub 2007 Dec 26. Nucleic Acids Res. 2008. PMID: 18160411 Free PMC article.
-
Using an RNA secondary structure partition function to determine confidence in base pairs predicted by free energy minimization.RNA. 2004 Aug;10(8):1178-90. doi: 10.1261/rna.7650904. RNA. 2004. PMID: 15272118 Free PMC article.
-
NMR structure of the A730 loop of the Neurospora VS ribozyme: insights into the formation of the active site.Nucleic Acids Res. 2011 May;39(10):4427-37. doi: 10.1093/nar/gkq1244. Epub 2011 Jan 25. Nucleic Acids Res. 2011. PMID: 21266483 Free PMC article.
-
Mg2+ Binding Promotes SLV as a Scaffold in Varkud Satellite Ribozyme SLI-SLV Kissing Loop Junction.Biophys J. 2017 Jul 25;113(2):313-320. doi: 10.1016/j.bpj.2017.06.008. Epub 2017 Jun 29. Biophys J. 2017. PMID: 28669407 Free PMC article.
-
A guanine nucleobase important for catalysis by the VS ribozyme.EMBO J. 2007 May 16;26(10):2489-500. doi: 10.1038/sj.emboj.7601698. Epub 2007 Apr 26. EMBO J. 2007. PMID: 17464286 Free PMC article.
References
-
- Doudna, J. A. & Cech, T. R. (2002) Nature 418, 221–228. - PubMed
-
- Murray, J. B., Dunham, C. M. & Scott, W. G. (2002) J. Mol. Biol. 315, 121–130. - PubMed
-
- Saville, B. J. & Collins, R. A. (1990) Cell 61, 685–696. - PubMed
-
- Guo, H. C. T., De Abreu, D. M., Tillier, E. R. M., Saville, B. J., Olive, J. E. & Collins, R. A. (1993) J. Mol. Biol. 232, 351–361. - PubMed
-
- Rastogi, T. & Collins, R. A. (1998) J. Mol. Biol. 277, 215–224. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

