Limitations of TaqMan PCR for detecting divergent viral pathogens illustrated by hepatitis A, B, C, and E viruses and human immunodeficiency virus
- PMID: 12791858
- PMCID: PMC156483
- DOI: 10.1128/JCM.41.6.2417-2427.2003
Limitations of TaqMan PCR for detecting divergent viral pathogens illustrated by hepatitis A, B, C, and E viruses and human immunodeficiency virus
Abstract
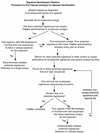
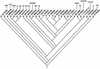
Recent events illustrate the imperative to rapidly and accurately detect and identify pathogens during disease outbreaks, whether they are natural or engineered. Particularly for our primary goal of detecting bioterrorist releases, detection techniques must be both species-wide (capable of detecting all known strains of a given species) and species specific. Due to classification restrictions on the publication of data for species that may pose a bioterror threat, we illustrate the challenges of finding such assays using five nonthreat organisms that are nevertheless of public health concern: human immunodeficiency virus (HIV) and four species of hepatitis viruses. Fluorogenic probe-based PCR assays (TaqMan; Perkin-Elmer Corp., Applied Biosystems, Foster City, Calif.) may be sensitive, fast methods for the identification of species in which the genome is conserved among strains, such as hepatitis A virus. For species such as HIV, however, the strains are highly divergent. We use computational methods to show that nine TaqMan primer and probe sequences, or signatures, are needed to ensure that all strains will be detected, but this is an unfeasible number, considering the cost of TaqMan probes. Strains of hepatitis B, C, and E viruses show intermediate divergence, so that two to three TaqMan signatures are required to detect all strains of each virus. We conclude that for species such as hepatitis A virus with high levels of sequence conservation among strains, signatures can be found computationally for detection by the TaqMan assay, which is a sensitive, rapid, and cost-effective method. However, for species such as HIV with substantial genetic divergence among strains, the TaqMan assay becomes unfeasible and alternative detection methods may be required. We compare the TaqMan assay with some of the alternative nucleic acid-based detection techniques of microarray, chip, and bead technologies in terms of sensitivity, speed, and cost.
Figures

References
-
- Garin, D., C. Peyrefitte, J. M. Crance, A. Le Faou, A. Jouan, and M. Bouloy. 2001. Highly sensitive TaqMan® PCR detection of Puumala hantavirus. Microbes Infect. 3:739-745. - PubMed
-
- Hadfield, T. L., M. Turell, M. P. Dempsey, J. David, and E. J. Park. 2001. Detection of West Nile virus in mosquitoes by RT-PCR. Mol. Cell. Probes 15:147-150. - PubMed
-
- Lipshutz, R. J., S. P. A. Fodor, T. R. Gingeras, and D. J. Lockhart. 1999. High density synthetic oligonucleotide arrays. Nat. Genet. 21:20-24. - PubMed
-
- Livak, K., J. Marmaro, and S. Flood. 1995. Guidelines for designing TaqMan fluorogenic probes for 5′ nuclease assays. Perkin-Elmer, Boston, Mass.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

