Subgenotype analysis of Cryptosporidium isolates from humans, cattle, and zoo ruminants in Portugal
- PMID: 12791920
- PMCID: PMC156540
- DOI: 10.1128/JCM.41.6.2744-2747.2003
Subgenotype analysis of Cryptosporidium isolates from humans, cattle, and zoo ruminants in Portugal
Abstract
Cryptosporidium parvum and Cryptosporidium hominis isolates from human immunodeficiency virus-infected patients, cattle, and wild ruminants were characterized by PCR and DNA sequencing analysis of the 60-kDa glycoprotein gene. Seven alleles were identified, three corresponding to C. hominis and four corresponding to C. parvum. One new allele was found (IId), and one (IIb) had only been found in Portugal. Isolates from cattle and wild ruminants clustered in two alleles. In contrast, human isolates clustered in seven alleles, showing extensive allelic diversity.
Figures

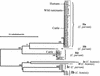
References
-
- Alves, M., O. Matos, and F. Antunes. 2001. Multilocus PCR-RFLP analysis of Cryptosporidium isolates from HIV-infected patients from Portugal. Ann. Trop. Med. Parasitol. 95:627-632. - PubMed
-
- Alves, M., O. Matos, F. Spano, and F. Antunes. 2000. PCR-RFLP analysis of Cryptosporidium parvum isolates from HIV-infected patients in Lisbon, Portugal. Ann. Trop. Med. Parasitol. 94:291-297. - PubMed
-
- Alves, M., O. Matos, I. P. Fonseca, E. Delgado, A. M. Lourenço, and F. Antunes. 2001. Multilocus genotyping of Cryptosporidium isolates from human HIV-infected and animal hosts. J. Eukarot. Microbiol. 2001 (Suppl.):17S-18S. - PubMed
Publication types
MeSH terms
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Medical

