Rapamycin treatment results in GATA factor-independent hyperphosphorylation of the proline utilization pathway activator in Saccharomyces cerevisiae
- PMID: 12796300
- PMCID: PMC161436
- DOI: 10.1128/EC.2.3.552-559.2003
Rapamycin treatment results in GATA factor-independent hyperphosphorylation of the proline utilization pathway activator in Saccharomyces cerevisiae
Abstract
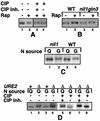
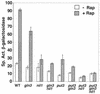
Treatment of Saccharomyces cerevisiae cells with the immunosuppressive drug rapamycin results in a variety of cellular changes in response to perceived nutrient deprivation. Among other effects, rapamycin treatment results in the nuclear localization of the global nitrogen activators Gln3p and Nil1p/Gat1p, which leads to expression of nitrogen assimilation genes. The proline utilization (Put) pathway genes were shown to be among the genes induced by rapamycin. Having previously shown that the Put pathway activator Put3p is differentially phosphorylated in response to the quality of the nitrogen source, we examined the phosphorylation status of Put3p after rapamycin treatment. Treatment with rapamycin resulted in the hyperphosphorylation of Put3p, which was independent of Gln3p, Nil1p, and Ure2p. The relative contributions of global nitrogen (Gln3p and Nil1p) and pathway-specific (Put3p) activators to rapamycin-induced expression of the target gene PUT1 were also examined. We found that Nil1p and Put3p, but not Gln3p, play major roles in rapamycin-induced PUT1 expression. Our findings show that perceived nitrogen deprivation triggered by rapamycin treatment and steady-state growth in nitrogen-derepressing conditions are associated with hyperphosphorylation of Put3p and increased PUT1 expression. Rapamycin treatment and nitrogen derepression may share some, but not all, regulatory elements, since Gln3p and Nil1p do not participate identically in both processes and are not required for hyperphosphorylation. A complex relationship exists among the global and pathway-specific regulators, depending on the nature and quality of the nitrogen source.
Figures

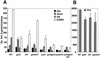

 , Put3p dimers;
, Put3p dimers;  , proline;
, proline;  , positive GATA factors;
, positive GATA factors;  , negative GATA factors.
, negative GATA factors.References
-
- Beck, T., and M. N. Hall. 1999. The TOR signalling pathway controls nuclear localization of nutrient-regulated transcription factors. Nature 402:689-692. - PubMed
-
- Bertram, P. G., J. H. Choi, J. Carvalho, W. Ai, C. Zeng, T.-F. Chan, and X. F. S. Zheng. 2000. Tripartite regulation of Gln3p by TOR, Ure2p and phosphatases. J. Biol. Chem. 275:35727-35733. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

