N-terminus of hMLH1 confers interaction of hMutLalpha and hMutLbeta with hMutSalpha
- PMID: 12799449
- PMCID: PMC162253
- DOI: 10.1093/nar/gkg420
N-terminus of hMLH1 confers interaction of hMutLalpha and hMutLbeta with hMutSalpha
Abstract
Mismatch repair is a highly conserved system that ensures replication fidelity by repairing mispairs after DNA synthesis. In humans, the two protein heterodimers hMutSalpha (hMSH2-hMSH6) and hMutLalpha (hMLH1-hPMS2) constitute the centre of the repair reaction. After recognising a DNA replication error, hMutSalpha recruits hMutLalpha, which then is thought to transduce the repair signal to the excision machinery. We have expressed an ATPase mutant of hMutLalpha as well as its individual subunits hMLH1 and hPMS2 and fragments of hMLH1, followed by examination of their interaction properties with hMutSalpha using a novel interaction assay. We show that, although the interaction requires ATP, hMutLalpha does not need to hydrolyse this nucleotide to join hMutSalpha on DNA, suggesting that ATP hydrolysis by hMutLalpha happens downstream of complex formation. The analysis of the individual subunits of hMutLalpha demonstrated that the hMutSalpha-hMutLalpha interaction is predominantly conferred by hMLH1. Further experiments revealed that only the N-terminus of hMLH1 confers this interaction. In contrast, only the C-terminus stabilised and co-immunoprecipitated hPMS2 when both proteins were co-expressed in 293T cells, indicating that dimerisation and stabilisation are mediated by the C-terminal part of hMLH1. We also examined another human homologue of bacterial MutL, hMutLbeta (hMLH1-hPMS1). We show that hMutLbeta interacts as efficiently with hMutSalpha as hMutLalpha, and that it predominantly binds to hMutSalpha via hMLH1 as well.
Figures








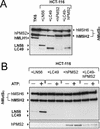


References
-
- Marti T.M., Kunz,C. and Fleck,O. (2002) DNA mismatch repair and mutation avoidance pathways. J. Cell. Physiol., 191, 28–41. - PubMed
-
- Modrich P. (1997) Strand-specific mismatch repair in mammalian cells. J. Biol. Chem., 272, 24727–24730. - PubMed
-
- Peltomaki P. (2001) DNA mismatch repair and cancer. Mutat. Res., 488, 77–85. - PubMed
-
- Lynch H.T. (1999) Hereditary nonpolyposis colorectal cancer (HNPCC). Cytogenet. Cell Genet., 86, 130–135. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

