Genomic transcriptional profiling of the developmental cycle of Chlamydia trachomatis
- PMID: 12815105
- PMCID: PMC166254
- DOI: 10.1073/pnas.1331135100
Genomic transcriptional profiling of the developmental cycle of Chlamydia trachomatis
Abstract
Chlamydia trachomatis is one of the most common bacterial pathogens and is the etiological agent of debilitating sexually transmitted and ocular diseases in humans. The organism is an obligate intracellular prokaryote characterized by a highly specialized biphasic developmental cycle. We have performed genomic transcriptional analysis of the chlamydial developmental cycle. This approach has led to the identification of a small subset of genes that control the primary (immediate-early genes) and secondary (late genes) differentiation stages of the cycle. Immediate-early gene products initiate bacterial metabolism and potentially modify the bacterial phagosome to escape fusion with lysosomes. One immediate early gene (CT147) is a homolog of the human early endosomal antigen-1 that is localized to the chlamydial phagosome; suggesting a functional role for CT147 in establishing the parasitophorous vacuole in a nonfusogenic pathway. Late gene products terminate bacterial cell division and constitute structural components and remodeling activities involved in the formation of the highly disulfide cross-linked outer-membrane complex that functions in attachment and invasion of new host cells. Many of the genes expressed during the immediate-early and late differentiation stages are Chlamydia-specific and have evolutionary origins in eukaryotic lineages.
Figures
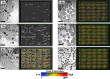



Comment in
-
Chlamydia uncloaked.Proc Natl Acad Sci U S A. 2003 Jul 8;100(14):8040-2. doi: 10.1073/pnas.1533181100. Epub 2003 Jun 30. Proc Natl Acad Sci U S A. 2003. PMID: 12835422 Free PMC article. No abstract available.
References
-
- Schachter, J. (1978) N. Engl. J. Med. 298 428-434. - PubMed
-
- Brunham, R. C., Binns, B., Guijon, F., Danforth, D., Kosseim, M. L., Rand, F., McDowell, J. & Rayner, E. (1988) J. Infect. Dis. 158 510-517. - PubMed
-
- Anttila, T., Saikku, P., Koskela, P., Bloigu, A., Dillner, J., Ikaheimo, I., Jellum, E., Lehtinen, M., Lenner, P., Hakulinen, T., et al. (2001) J. Am. Med. Assoc. 285 47-51. - PubMed
-
- Chesson, H. W. & Pinkerton, S. D. (2000) J. Acquired Immune Defic. Syndr. 24 48-56. - PubMed
-
- Mbizvo, E. M., Msuya, S. E., Stray-Pedersen, B., Sundby, J., Chirenje, M. Z. & Hussain, A. (2001) Int. J. STD AIDS 12 524-531. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

