Phenotypic and molecular characterization of tetracycline- and erythromycin-resistant strains of Streptococcus pneumoniae
- PMID: 12821474
- PMCID: PMC161878
- DOI: 10.1128/AAC.47.7.2236-2241.2003
Phenotypic and molecular characterization of tetracycline- and erythromycin-resistant strains of Streptococcus pneumoniae
Abstract
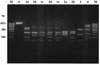
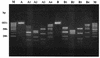
Sixty-five clinical isolates of Streptococcus pneumoniae, all collected in Italy between 1999 and 2002 and resistant to both tetracycline (MIC, >or=8 microg/ml) and erythromycin (MIC, >or=1 microg/ml), were investigated. Of these strains, 11% were penicillin resistant and 23% were penicillin intermediate. With the use of the erythromycin-clindamycin-rokitamycin triple-disk test, 14 strains were assigned to the constitutive (cMLS) phenotype of macrolide resistance, 44 were assigned to the partially inducible (iMcLS) phenotype, 1 was assigned to the inducible (iMLS) phenotype, and 6 were assigned to the efflux-mediated (M) phenotype. In PCR assays, 64 of the 65 strains were positive for the tetracycline resistance gene tet(M), the exception being the one M isolate susceptible to kanamycin, whereas tet(K), tet(L), and tet(O) were never found. All cMLS, iMcLS, and iMLS isolates had the erythromycin resistance gene erm(B), and all M phenotype isolates had the mef(A) or mef(E) gene. No isolate had the erm(A) gene. The int-Tn gene, encoding the integrase of the Tn916-Tn1545 family of conjugative transposons, was detected in 62 of the 65 test strains. Typing assays showed the strains to be to a great extent unrelated. Of 16 different serotypes detected, the most numerous were 23F (n = 13), 19A (n = 10), 19F (n = 9), 6B (n = 8), and 14 (n = 6). Of 49 different pulsed-field gel electrophoresis types identified, the majority (n = 39) were represented by a single isolate, while the most numerous type included five isolates. By high-resolution restriction analysis of PCR amplicons with four endonucleases, the tet(M) loci from the 64 tet(M)-positive pneumococci were classified into seven distinct restriction types. Overall, a Tn1545-like transposon could reasonably account for tetracycline and erythromycin resistance in the vast majority of the pneumococci of cMLS, iMcLS, and iMLS phenotypes, whereas a Tn916-like transposon could account for tetracycline resistance in most M phenotype strains.
Figures
Similar articles
-
Genetic elements carrying erm(B) in Streptococcus pyogenes and association with tet(M) tetracycline resistance gene.Antimicrob Agents Chemother. 2007 Apr;51(4):1209-16. doi: 10.1128/AAC.01484-06. Epub 2007 Jan 29. Antimicrob Agents Chemother. 2007. PMID: 17261630 Free PMC article.
-
Serotypes, Clones, and Mechanisms of Resistance of Erythromycin-Resistant Streptococcus pneumoniae Isolates Collected in Spain.Antimicrob Agents Chemother. 2007 Sep;51(9):3240-6. doi: 10.1128/AAC.00157-07. Epub 2007 Jul 2. Antimicrob Agents Chemother. 2007. PMID: 17606677 Free PMC article.
-
Differentiation of resistance phenotypes among erythromycin-resistant Pneumococci.J Clin Microbiol. 2001 Apr;39(4):1311-5. doi: 10.1128/JCM.39.4.1311-1315.2001. J Clin Microbiol. 2001. PMID: 11283047 Free PMC article.
-
erm(B)-carrying elements in tetracycline-resistant pneumococci and correspondence between Tn1545 and Tn6003.Antimicrob Agents Chemother. 2008 Apr;52(4):1285-90. doi: 10.1128/AAC.01457-07. Epub 2008 Feb 19. Antimicrob Agents Chemother. 2008. PMID: 18285489 Free PMC article.
-
Phenotypes and genotypes of erythromycin-resistant Streptococcus pyogenes strains in Italy and heterogeneity of inducibly resistant strains.Antimicrob Agents Chemother. 1999 Aug;43(8):1935-40. doi: 10.1128/AAC.43.8.1935. Antimicrob Agents Chemother. 1999. PMID: 10428916 Free PMC article.
Cited by
-
Antimicrobial susceptibility of invasive Streptococcus pneumoniae isolates in Portugal over an 11-year period.Antimicrob Agents Chemother. 2006 Jun;50(6):2098-105. doi: 10.1128/AAC.00198-06. Antimicrob Agents Chemother. 2006. PMID: 16723571 Free PMC article.
-
emm Types, virulence factors, and antibiotic resistance of invasive Streptococcus pyogenes isolates from Italy: What has changed in 11 years?J Clin Microbiol. 2007 Jul;45(7):2249-56. doi: 10.1128/JCM.00513-07. Epub 2007 May 9. J Clin Microbiol. 2007. PMID: 17494723 Free PMC article.
-
Prevalence of antibiotic resistance genes of wastewater and surface water in livestock farms of Jiangsu Province, China.Environ Sci Pollut Res Int. 2015 Sep;22(18):13950-9. doi: 10.1007/s11356-015-4636-y. Epub 2015 May 8. Environ Sci Pollut Res Int. 2015. PMID: 25948386
-
Effect of presumptive co-trimoxazole prophylaxis on pneumococcal colonization rates, seroepidemiology and antibiotic resistance in Zambian infants: a longitudinal cohort study.Bull World Health Organ. 2008 Dec;86(12):929-38. doi: 10.2471/blt.07.049668. Bull World Health Organ. 2008. PMID: 19142293 Free PMC article.
-
A comparison of various feature extraction and machine learning methods for antimicrobial resistance prediction in streptococcus pneumoniae.Front Antibiot. 2023 Mar 24;2:1126468. doi: 10.3389/frabi.2023.1126468. eCollection 2023. Front Antibiot. 2023. PMID: 39816648 Free PMC article.
References
-
- Betriu, C., M. Redondo, M. L. Palau, A. Sánchez, M. Gómez, E. Culebras, A. Boloix, and J. J. Picazo. 2000. Comparative in vitro activities of linezolid, quinupristin-dalfopristin, moxifloxacin, and trovafloxacin against erythromycin-susceptible and -resistant streptococci. Antimicrob. Agents Chemother. 44:1838-1841. - PMC - PubMed
-
- Clancy, J., J. Petitpas, F. Dib-Hajj, W. Yuan, M. Cronan, A. V. Kamath, J. Bergeron, and J. A. Retsema. 1996. Molecular cloning and functional analysis of a novel macrolide-resistance determinant, mefA, from Streptococcus pyogenes. Mol. Microbiol. 22:867-879. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

