Pfold: RNA secondary structure prediction using stochastic context-free grammars
- PMID: 12824339
- PMCID: PMC169020
- DOI: 10.1093/nar/gkg614
Pfold: RNA secondary structure prediction using stochastic context-free grammars
Abstract
RNA secondary structures are important in many biological processes and efficient structure prediction can give vital directions for experimental investigations. Many available programs for RNA secondary structure prediction only use a single sequence at a time. This may be sufficient in some applications, but often it is possible to obtain related RNA sequences with conserved secondary structure. These should be included in structural analyses to give improved results. This work presents a practical way of predicting RNA secondary structure that is especially useful when related sequences can be obtained. The method improves a previous algorithm based on an explicit evolutionary model and a probabilistic model of structures. Predictions can be done on a web server at http://www.daimi.au.dk/~compbio/pfold.
Figures

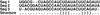
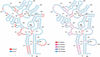
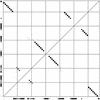
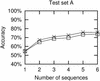


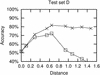
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

