Comprehensive quantitative analyses of the effects of promoter sequence elements on mRNA transcription
- PMID: 12824429
- PMCID: PMC168999
- DOI: 10.1093/nar/gkg593
Comprehensive quantitative analyses of the effects of promoter sequence elements on mRNA transcription
Abstract
We have generated a WWW interface for automated comprehensive analyses of promoter regulatory motifs and the effect they exert on mRNA expression profiles. The server provides a wide spectrum of analysis tools that allow de novo discovery of regulatory motifs, along with refinement and in-depth investigation of fully or partially characterized motifs. The presented discovery and analysis tools are fundamentally different from existing tools in their basic rational, statistical background and specificity and sensitivity towards true regulatory elements. We thus anticipate that the service will be of great importance to the experimental and computational biology communities alike. The motif discovery and diagnosis workbench is available at http://longitude.weizmann.ac.il/rMotif/.
Figures

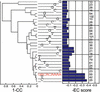
References
-
- Werner T. (2002) Finding and decrypting of promoters contributes to the elucidation of gene function. In Silico Biol., 2, 249–255. - PubMed
-
- Tavazoie S., Hughes,J.D., Campbell,M.J., Cho,R.J. and Church,G.M. (1999) Systematic determination of genetic network architecture. Nature Genet., 22, 281–285. - PubMed

