Predicting the topology of transmembrane helical proteins using mean burial propensity and a hidden-Markov-model-based method
- PMID: 12824500
- PMCID: PMC2323935
- DOI: 10.1110/ps.0305103
Predicting the topology of transmembrane helical proteins using mean burial propensity and a hidden-Markov-model-based method
Abstract
Helices in membrane spanning regions are more tightly packed than the helices in soluble proteins. Thus, we introduce a method that uses a simple scale of burial propensity and a new algorithm to predict transmembrane helical (TMH) segments and a positive-inside rule to predict amino-terminal orientation. The method (the topology predictor of transmembrane helical proteins using mean burial propensity [THUMBUP]) correctly predicted the topology of 55 of 73 proteins (or 75%) with known three-dimensional structures (the 3D helix database). This level of accuracy can be reached by MEMSAT 1.8 (a 200-parameter model-recognition method) and a new HMM-based method (a 111-parameter hidden Markov model, UMDHMM(TMHP)) if they were retrained with the 73-protein database. Thus, a method based on a physiochemical property can provide topology prediction as accurate as those methods based on more complicated statistical models and learning algorithms for the proteins with accurately known structures. Commonly used HMM-based methods and MEMSAT 1.8 were trained with a combination of the partial 3D helix database and a 1D helix database of TMH proteins in which topology information were obtained by gene fusion and other experimental techniques. These methods provide a significantly poorer prediction for the topology of TMH proteins in the 3D helix database. This suggests that the 1D helix database, because of its inaccuracy, should be avoided as either a training or testing database. A Web server of THUMBUP and UMDHMM(TMHP) is established for academic users at http://www.smbs.buffalo.edu/phys_bio/service.htm. The 3D helix database is also available from the same Web site.
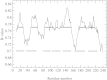
Figures
Similar articles
-
Web-based toolkits for topology prediction of transmembrane helical proteins, fold recognition, structure and binding scoring, folding-kinetics analysis and comparative analysis of domain combinations.Nucleic Acids Res. 2005 Jul 1;33(Web Server issue):W193-7. doi: 10.1093/nar/gki360. Nucleic Acids Res. 2005. PMID: 15980453 Free PMC article.
-
A hidden Markov model with molecular mechanics energy-scoring function for transmembrane helix prediction.Comput Biol Chem. 2004 Oct;28(4):265-74. doi: 10.1016/j.compbiolchem.2004.07.002. Comput Biol Chem. 2004. PMID: 15548453
-
Combined prediction of transmembrane topology and signal peptide of beta-barrel proteins: using a hidden Markov model and genetic algorithms.Comput Biol Med. 2010 Jul;40(7):621-8. doi: 10.1016/j.compbiomed.2010.04.006. Epub 2010 May 21. Comput Biol Med. 2010. PMID: 20488436
-
Topology prediction of helical transmembrane proteins: how far have we reached?Curr Protein Pept Sci. 2010 Nov;11(7):550-61. doi: 10.2174/138920310794109184. Curr Protein Pept Sci. 2010. PMID: 20887261 Review.
-
Topology of membrane proteins-predictions, limitations and variations.Curr Opin Struct Biol. 2018 Jun;50:9-17. doi: 10.1016/j.sbi.2017.10.003. Epub 2017 Nov 5. Curr Opin Struct Biol. 2018. PMID: 29100082 Review.
Cited by
-
A complementary bioinformatics approach to identify potential plant cell wall glycosyltransferase-encoding genes.Plant Physiol. 2004 Sep;136(1):2609-20. doi: 10.1104/pp.104.042978. Epub 2004 Aug 27. Plant Physiol. 2004. PMID: 15333752 Free PMC article.
-
Algorithms for incorporating prior topological information in HMMs: application to transmembrane proteins.BMC Bioinformatics. 2006 Apr 5;7:189. doi: 10.1186/1471-2105-7-189. BMC Bioinformatics. 2006. PMID: 16597327 Free PMC article.
-
Bias-Exchange Metadynamics Simulation of Membrane Permeation of 20 Amino Acids.Int J Mol Sci. 2018 Mar 16;19(3):885. doi: 10.3390/ijms19030885. Int J Mol Sci. 2018. PMID: 29547563 Free PMC article.
-
The vegetative vacuole proteome of Arabidopsis thaliana reveals predicted and unexpected proteins.Plant Cell. 2004 Dec;16(12):3285-303. doi: 10.1105/tpc.104.027078. Epub 2004 Nov 11. Plant Cell. 2004. PMID: 15539469 Free PMC article.
-
MemBrain: improving the accuracy of predicting transmembrane helices.PLoS One. 2008 Jun 11;3(6):e2399. doi: 10.1371/journal.pone.0002399. PLoS One. 2008. PMID: 18545655 Free PMC article.
References
-
- Argos, P., Rao, J.K., and Hargrave, P.A. 1982. Structural prediction of membrane-bound proteins. Eur. J. Biochem. 128 565–575. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

