Characterization of adapt33, a stress-inducible riboregulator
- PMID: 12837039
- PMCID: PMC5991141
- DOI: 10.3727/000000003108748982
Characterization of adapt33, a stress-inducible riboregulator
Abstract
We have identified adapt33 as a multiple stress-responsive gene that is induced under conditions of a cytoprotective "adaptive response." adapt33 RNA does not contain any appreciable open reading frame nor produce a protein product and is therefore classified as a stress-inducible riboregulator. Although a number of oxidant stress-modulated, protein-encoding genes have been reported and characterized, very few stress-inducible riboregulator RNAs are known. Here we extend previous studies toward understanding the underlying regulation of expression and function of this rare mammalian riboregulator. mRNA stability and transcription studies determined that adapt33 induction by hydrogen peroxide is at the mRNA stability level, and that adapt33 has a very short half-life. Surprisingly, adapt33 mRNA also exhibits altered electrophoretic migration in response to both hydrogen peroxide and cis-platinum treatment. Although no transcriptional modulation in response to hydrogen peroxide was observed, fusion promoter constructs revealed that adapt33 has an unusually strong promoter that is active in both hamster and human cells. Analysis of expression following the stimulation of apoptosis with hydrogen peroxide and staurosporine revealed a strong correlation with apoptosis, suggesting a possible novel, noncoding RNA component of the apoptotic mechanism. We conclude that adapt33 is a stress-inducible, apoptosis-associated RNA with unique structural and gene promoter characteristics.
Figures




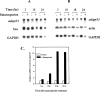
Similar articles
-
adapt33, a novel oxidant-inducible RNA from hamster HA-1 cells.Arch Biochem Biophys. 1996 Aug 15;332(2):255-60. doi: 10.1006/abbi.1996.0340. Arch Biochem Biophys. 1996. PMID: 8806733
-
Modulation of a cardiogenic shock inducible RNA by chemical stress: adapt73/PigHep3.Surgery. 1997 May;121(5):581-7. doi: 10.1016/s0039-6060(97)90115-x. Surgery. 1997. PMID: 9142159
-
Apurinic endonuclease (Ref-1) is induced in mammalian cells by oxidative stress and involved in clastogenic adaptation.Cancer Res. 1998 Oct 1;58(19):4410-6. Cancer Res. 1998. PMID: 9766671
-
Hydrogen peroxide induces the expression of adapt15, a novel RNA associated with polysomes in hamster HA-1 cells.Arch Biochem Biophys. 1996 Jan 15;325(2):256-64. doi: 10.1006/abbi.1996.0032. Arch Biochem Biophys. 1996. PMID: 8561505
-
Hamster adapt78 mRNA is a Down syndrome critical region homologue that is inducible by oxidative stress.Arch Biochem Biophys. 1997 Jun 1;342(1):6-12. doi: 10.1006/abbi.1997.0109. Arch Biochem Biophys. 1997. PMID: 9185608
Cited by
-
Genome-wide identification and characterization of long non-coding RNAs involved in the early somatic embryogenesis in Dimocarpus longan Lour.BMC Genomics. 2018 Nov 6;19(1):805. doi: 10.1186/s12864-018-5158-z. BMC Genomics. 2018. PMID: 30400813 Free PMC article.
-
Divergent regulation of lncRNA expression by ischemia in adult and aging mice.Geroscience. 2022 Feb;44(1):429-445. doi: 10.1007/s11357-021-00460-9. Epub 2021 Oct 26. Geroscience. 2022. PMID: 34697716 Free PMC article.
-
Annotation of long non-coding RNAs expressed in collaborative cross founder mice in response to respiratory virus infection reveals a new class of interferon-stimulated transcripts.RNA Biol. 2014;11(7):875-90. doi: 10.4161/rna.29442. Epub 2014 Jun 12. RNA Biol. 2014. PMID: 24922324 Free PMC article.
-
Nonsense-mediated decay factor SMG7 sensitizes cells to TNFα-induced apoptosis via CYLD tumor suppressor and the noncoding oncogene Pvt1.Mol Oncol. 2020 Oct;14(10):2420-2435. doi: 10.1002/1878-0261.12754. Epub 2020 Jul 13. Mol Oncol. 2020. PMID: 32602581 Free PMC article.
-
lncRNA 5430416N02Rik Promotes the Proliferation of Mouse Embryonic Stem Cells by Activating Mid1 Expression through 3D Chromatin Architecture.Stem Cell Reports. 2020 Mar 10;14(3):493-505. doi: 10.1016/j.stemcr.2020.02.002. Stem Cell Reports. 2020. PMID: 32160522 Free PMC article.
References
-
- Bargonetti J.; Manfredi J. J. Multiple roles of the tumor suppressor p53. Curr. Opin. Oncol. 14:86–91; 2002. - PubMed
-
- Brockdorff N. A.; Ashworth G. F.; Kay V. M.; McCabe D. P.; Norris P. J.; Cooper S. S.; Rastan S. The product of the mouse Xist gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell 71:515–526; 1992. - PubMed
-
- Brunkow M. E.; Tilghman S. M. Ectopic expression of the H19 gene in mice causes prenatal lethality. Genes Dev. 5:1092–1101; 1991. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
