Structure of a coenzyme A pyrophosphatase from Deinococcus radiodurans: a member of the Nudix family
- PMID: 12837785
- PMCID: PMC164880
- DOI: 10.1128/JB.185.14.4110-4118.2003
Structure of a coenzyme A pyrophosphatase from Deinococcus radiodurans: a member of the Nudix family
Abstract
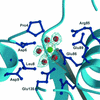
Gene Dr1184 from Deinococcus radiodurans codes for a Nudix enzyme (DR-CoAse) that hydrolyzes the pyrophosphate moiety of coenzyme A (CoA). Nudix enzymes with the same specificity have been found in yeast, humans, and mice. The three-dimensional structure of DR-CoAse, the first of a Nudix hydrolase with this specificity, reveals that this enzyme contains, in addition to the fold observed in other Nudix enzymes, insertions that are characteristic of a CoA-hydrolyzing Nudix subfamily. The structure of the complex of the enzyme with Mg(2+), its activating cation, reveals the position of the catalytic site. A helix, part of the N-terminal insertion, partially occludes the binding site and has to change its position to permit substrate binding. Comparison of the structure of DR-CoAse to those of other Nudix enzymes, together with the location in the structure of the sequence characteristic of CoAses, suggests a mode of binding of the substrate to the enzyme that is compatible with all available data.
Figures




Similar articles
-
Structure of an N-terminally truncated Nudix hydrolase DR2204 from Deinococcus radiodurans.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009 Nov 1;65(Pt 11):1083-7. doi: 10.1107/S1744309109037191. Epub 2009 Oct 13. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009. PMID: 19923723 Free PMC article.
-
Functional and structural characterization of DR_0079 from Deinococcus radiodurans, a novel Nudix hydrolase with a preference for cytosine (deoxy)ribonucleoside 5'-Di- and triphosphates.Biochemistry. 2008 Jun 24;47(25):6571-82. doi: 10.1021/bi800099d. Biochemistry. 2008. PMID: 18512963 Free PMC article.
-
Solution structure of hypothetical Nudix hydrolase DR0079 from extremely radiation-resistant Deinococcus radiodurans bacterium.Proteins. 2004 Jul 1;56(1):28-39. doi: 10.1002/prot.20082. Proteins. 2004. PMID: 15162484
-
Structures and mechanisms of Nudix hydrolases.Arch Biochem Biophys. 2005 Jan 1;433(1):129-43. doi: 10.1016/j.abb.2004.08.017. Arch Biochem Biophys. 2005. PMID: 15581572 Review.
-
Nudix proteins affecting microbial pathogenesis.Microbiology (Reading). 2020 Dec;166(12):1110-1114. doi: 10.1099/mic.0.000993. Microbiology (Reading). 2020. PMID: 33253082 Review.
Cited by
-
Structural and Enzymatic Characterization of a Nucleoside Diphosphate Sugar Hydrolase from Bdellovibrio bacteriovorus.PLoS One. 2015 Nov 2;10(11):e0141716. doi: 10.1371/journal.pone.0141716. eCollection 2015. PLoS One. 2015. PMID: 26524597 Free PMC article.
-
Structural analyses of NudT16-ADP-ribose complexes direct rational design of mutants with improved processing of poly(ADP-ribosyl)ated proteins.Sci Rep. 2019 Apr 11;9(1):5940. doi: 10.1038/s41598-019-39491-w. Sci Rep. 2019. PMID: 30976021 Free PMC article.
-
Biosynthesis of Pantothenic Acid and Coenzyme A.EcoSal Plus. 2007 Apr;2(2):10.1128/ecosalplus.3.6.3.4. doi: 10.1128/ecosalplus.3.6.3.4. EcoSal Plus. 2007. PMID: 26443589 Free PMC article.
-
Identification of Nudix hydrolase family members with an antimutator role in Mycobacterium tuberculosis and Mycobacterium smegmatis.J Bacteriol. 2006 Apr;188(8):3159-61. doi: 10.1128/JB.188.8.3159-3161.2006. J Bacteriol. 2006. PMID: 16585780 Free PMC article.
-
Structure of an N-terminally truncated Nudix hydrolase DR2204 from Deinococcus radiodurans.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009 Nov 1;65(Pt 11):1083-7. doi: 10.1107/S1744309109037191. Epub 2009 Oct 13. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009. PMID: 19923723 Free PMC article.
References
-
- Abdelghany, H. M., L. Gasmi, J. L. Cartwright, S. Bailey, J. B. Rafferty, and A. G. McLennan. 2001. Cloning, characterisation and crystallisation of a diadenosine 5′,5‴-P(1),P(4)-tetraphosphate pyrophosphohydrolase from Caenorhabditis elegans. Biochim. Biophys. Acta 1550:27-36. - PubMed
-
- Abeygunawardana, C., D. J. Weber, A. G. Gittis, D. N. Frick, J. Lin, A. F. Miller, M. J. Bessman, and A. S. Mildvan. 1995. Solution structure of the MutT enzyme, a nucleoside triphosphate pyrophosphohydrolase. Biochemistry 34:14997-15005. - PubMed
-
- Bessman, M. J., D. N. Frick, and S. F. O'Handley. 1996. The MutT proteins or “Nudix” hydrolases, a family of versatile, widely distributed, “housecleaning” enzymes. J. Biol. Chem. 271:25059-25062. - PubMed
-
- Bhatnagar, S. K., L. C. Bullions, and M. J. Bessman. 1991. Characterization of the mutT nucleoside triphosphatase of Escherichia coli. J. Biol. Chem. 266:9050-9054. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

