Digestion pattern of reovirus outer capsid protein sigma3 determined by mass spectrometry
- PMID: 12842619
- PMCID: PMC7202455
- DOI: 10.1016/s0042-6822(03)00154-5
Digestion pattern of reovirus outer capsid protein sigma3 determined by mass spectrometry
Abstract
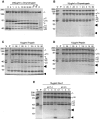
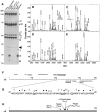
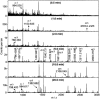
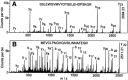
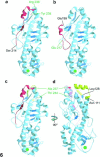
Reovirus is an enteric virus comprising eight structural proteins that form a double-layered capsid. During reovirus entry into cells, the outermost capsid layer (composed of proteins sigma3 and mu1C) is proteolytically processed to generate an infectious subviral particle (ISVP) that is subsequently uncoated to produce the transcriptionally active core particle. Kinetic studies suggest that protein sigma3 is rapidly removed from virus particles and then protein mu1C is cleaved. Initial cleavage of mu1C has been well described and generates an amino (N)-terminal delta peptide and a carboxyl (C)-terminal phi peptide. However, cleavage and removal of sigma3 is an extremely rapid event that has not been well defined. We have treated purified reovirus serotype 1 Lang virions with a variety of endoproteases. Time-course digestions with chymotrypsin, Glu-C, pepsin, and trypsin resulted in the initial generation of two peptides that were resolved in SDS-PAGE and analyzed by in-gel tryptic digestion and MALDI-Qq-TOFMS. Most tested proteases cut sigma3 within a "hypersensitive" region between amino acids 217 and 238. In addition, to gain a better understanding of the sequence of subsequent proteolytic events that result in generation of reovirus subviral particles, time-course digestions of purified particles were performed under physiologic salt conditions and released peptide fragments ranging from 500 to 5000 Da were directly analyzed by MALDI-Qq-TOFMS. Trypsin digestion initially released a peptide that corresponded to the C-terminus of mu1C, followed by a peptide that corresponded to amino acids 214-236 of the sigma3 protein. Other regions of mu1C were not observed until protein sigma3 was completely digested. Similar experiments with Glu-C indicated the hypersensitive region of sigma3 was cut first when virions were treated at pH values of 4.5 or 7.4, but treatment of virions with pepsin at pH 3.0 released different sigma3 peptides, suggesting acid-induced conformational changes in this outer capsid protein. These studies also revealed that the N-terminus of sigma3 is acetylated.
Figures


References
-
- Antczak J.B., Joklik W.K. Reovirus genome segment assortment into progeny genomes studied by the use of monoclonal antibodies directed against reovirus proteins. Virology. 1992;187:760–776. - PubMed
-
- Berger E.M., Cox G., Weber L., Kenney J.S. Actin acetylation in Drosophila tissue culture cells. Biochem. Genet. 1981;19:321–331. - PubMed
-
- Borsa J., Morash B.D., Sargent M.D., Copps T.P., Lievaart P.A., Szekely J.G. Two modes of entry of reovirus particles into L cells. J. Gen. Virol. 1979;45:161–170. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

