Molecular mechanism of signal sequence orientation in the endoplasmic reticulum
- PMID: 12853479
- PMCID: PMC165631
- DOI: 10.1093/emboj/cdg361
Molecular mechanism of signal sequence orientation in the endoplasmic reticulum
Abstract
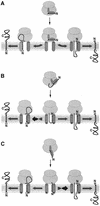
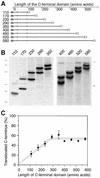
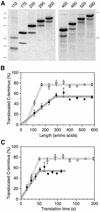
We have analyzed in vivo how model signal sequences are inserted and oriented in the membrane during cotranslational integration into the endoplasmic reticulum. The results are incompatible with the current models of retention of positive flanking charges or loop insertion of the polypeptide into the translocon. Instead they indicate that these N-terminal signals initially insert head-on with a cytoplasmic C-terminus before they invert their orientation to translocate the C-terminus. The rate of inversion increases with more positive N-terminal charge and is reduced with increasing hydrophobicity of the signal. Inversion may proceed for up to approximately 50 s, when it is terminated by a signal-independent process. These findings provide a mechanism for the topogenic effects of flanking charges as well as of signal hydrophobicity.
Figures


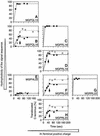
References
-
- Andrews D.W., Young,J.C., Mirels,L.F. and Czarnota,G.J. (1992) The role of the N-region in signal sequence and signal–anchor function. J. Biol. Chem., 267, 7761–7769. - PubMed
-
- Batey R.T., Rambo,R.P., Lucast,L., Rha,B. and Doudna,J.A. (2000) Crystal structure of the ribonucleoprotein core of the signal recognition particle. Science, 287, 1232–1239. - PubMed
-
- Beltzer J.P., Fiedler,K., Fuhrer,C., Geffen,I., Handschin,C., Wessels,H.P. and Spiess,M. (1991) Charged residues are major determinants of the transmembrane orientation of a signal–anchor sequence. J. Biol. Chem., 266, 973–978. - PubMed
-
- Chuck S.L. and Lingappa,V.R. (1992) Pause transfer: a topogenic sequence in apolipoprotein B mediates stopping and restarting of translocation. Cell, 68, 9–21. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

