Processing of DNA lesions by archaeal DNA polymerases from Sulfolobus solfataricus
- PMID: 12853619
- PMCID: PMC165962
- DOI: 10.1093/nar/gkg447
Processing of DNA lesions by archaeal DNA polymerases from Sulfolobus solfataricus
Abstract
Spontaneous damage to DNA as a result of deamination, oxidation and depurination is greatly accelerated at high temperatures. Hyperthermophilic microorganisms constantly exposed to temperatures exceeding 80 degrees C are endowed with powerful DNA repair mechanisms to maintain genome stability. Of particular interest is the processing of DNA lesions during replication, which can result in fixed mutations. The hyperthermophilic crenarchaeon Sulfolobus solfataricus has two functional DNA polymerases, PolB1 and PolY1. We have found that the replicative DNA polymerase PolB1 specifically recognizes the presence of the deaminated bases hypoxanthine and uracil in the template by stalling DNA polymerization 3-4 bases upstream of these lesions and strongly associates with oligonucleotides containing them. PolB1 also stops at 8-oxoguanine and is unable to bypass an abasic site in the template. PolY1 belongs to the family of lesion bypass DNA polymerases and readily bypasses hypoxanthine, uracil and 8-oxoguanine, but not an abasic site, in the template. The specific recognition of deaminated bases by PolB1 may represent an initial step in their repair while PolY1 may be involved in damage tolerance at the replication fork. Additionally, we reveal that the deaminated bases can be introduced into DNA enzymatically, since both PolB1 and PolY1 are able to incorporate the aberrant DNA precursors dUTP and dITP.
Figures



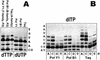


Similar articles
-
Kinetic basis for the differing response to an oxidative lesion by a replicative and a lesion bypass DNA polymerase from Sulfolobus solfataricus.Biochemistry. 2012 Apr 24;51(16):3485-96. doi: 10.1021/bi300246r. Epub 2012 Apr 10. Biochemistry. 2012. PMID: 22471521
-
Modulation of hyperthermophilic DNA polymerase activity by archaeal chromatin proteins.J Biol Chem. 2004 Jan 2;279(1):127-32. doi: 10.1074/jbc.M309860200. Epub 2003 Oct 16. J Biol Chem. 2004. PMID: 14563841
-
Cleavage of double-stranded DNA by the intrinsic 3'-5' exonuclease activity of DNA polymerase B1 from the hyperthermophilic archaeon Sulfolobus solfataricus at high temperature.FEMS Microbiol Lett. 2004 Feb 9;231(1):111-7. doi: 10.1016/S0378-1097(03)00932-7. FEMS Microbiol Lett. 2004. PMID: 14769474
-
Recognition of deaminated bases by archaeal family-B DNA polymerases.Biochem Soc Trans. 2009 Feb;37(Pt 1):65-8. doi: 10.1042/BST0370065. Biochem Soc Trans. 2009. PMID: 19143603 Review.
-
Recent insight into the kinetic mechanisms and conformational dynamics of Y-Family DNA polymerases.Biochemistry. 2014 May 6;53(17):2804-14. doi: 10.1021/bi5000405. Epub 2014 Apr 23. Biochemistry. 2014. PMID: 24716482 Free PMC article. Review.
Cited by
-
Enzymatic Switching Between Archaeal DNA Polymerases Facilitates Abasic Site Bypass.Front Microbiol. 2021 Dec 20;12:802670. doi: 10.3389/fmicb.2021.802670. eCollection 2021. Front Microbiol. 2021. PMID: 34987494 Free PMC article.
-
A Unique B-Family DNA Polymerase Facilitating Error-Prone DNA Damage Tolerance in Crenarchaeota.Front Microbiol. 2020 Jul 23;11:1585. doi: 10.3389/fmicb.2020.01585. eCollection 2020. Front Microbiol. 2020. PMID: 32793138 Free PMC article.
-
Biochemical evidence of a physical interaction between Sulfolobus solfataricus B-family and Y-family DNA polymerases.Extremophiles. 2007 Mar;11(2):277-82. doi: 10.1007/s00792-006-0038-x. Epub 2006 Nov 3. Extremophiles. 2007. PMID: 17082970
-
Accurate DNA synthesis by Sulfolobus solfataricus DNA polymerase B1 at high temperature.Extremophiles. 2010 Jan;14(1):107-17. doi: 10.1007/s00792-009-0292-9. Epub 2009 Dec 11. Extremophiles. 2010. PMID: 20012453
-
The 3'-5' proofreading exonuclease of archaeal family-B DNA polymerase hinders the copying of template strand deaminated bases.Nucleic Acids Res. 2009 Dec;37(22):7603-11. doi: 10.1093/nar/gkp800. Nucleic Acids Res. 2009. PMID: 19783818 Free PMC article.
References
-
- Fogg M.J., Pearl,L.H. and Connolly,B.A. (2002) Structural basis for uracil recognition by archaeal family B DNA polymerases. Nature Struct. Biol., 9, 922–927. - PubMed
-
- Kulaeva O.I., Koonin,E.V., McDonald,J.P., Randall,S.K., Rabinovich,N., Connaughton,J.F., Levine,A.S. and Woodgate,R. (1996) Identification of a DinB/UmuC homolog in the archeon Sulfolobus solfataricus. Mutat. Res., 357, 245–253. - PubMed
-
- Gruz P., Pisani,F.M., Shimizu,M., Yamada,M., Hayashi,I., Morikawa,K. and Nohmi,T. (2001) Synthetic activity of Sso DNA polymerase Y1, an archaeal DinB-like DNA polymerase, is stimulated by processivity factors proliferating cell nuclear antigen and replication factor C. J. Biol. Chem., 276, 47394–47401. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

