Selective interactions of human kin17 and RPA proteins with chromatin and the nuclear matrix in a DNA damage- and cell cycle-regulated manner
- PMID: 12853634
- PMCID: PMC165974
- DOI: 10.1093/nar/gkg459
Selective interactions of human kin17 and RPA proteins with chromatin and the nuclear matrix in a DNA damage- and cell cycle-regulated manner
Abstract
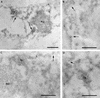
Several proteins involved in DNA synthesis are part of the so-called 'replication factories' that are anchored on non-chromatin nuclear structures. We report here that human kin17, a nuclear stress-activated protein, associates with both chromatin and non-chromatin nuclear structures in a cell cycle- and DNA damage-dependent manner. After L-mimosine block and withdrawal we observed that kin17 protein was recruited in the nucleus during re-entry and progression through S phase. These results are consistent with a role of kin17 protein in DNA replication. About 50% of the total amount of kin17 protein was detected on nuclear structures and could not be released by detergents. Furthermore, the amount of kin17 protein greatly increased in both G(1)/S and S phase-arrested cells in fractions containing proteins anchored to nuclear structures. The detection of kin17 protein showed for the first time its preferential assembly within non-chromatin nuclear structures in G(1)/S and S phase-arrested cells, while the association with these structures was found to be less stable in the G(2)/M phase, as judged by fractionation of human cells and immunostaining. In asynchronous growing cells, kin17 protein interacted with both chromatin DNA and non-chromatin nuclear structures, while in S phase-arrested cells it interacted mostly with non-chromatin nuclear structures, as judged by DNase I treatment and in vivo UV cross-linking. In the presence of DNA damage in S phase cells, the distribution of kin17 protein became mainly associated with chromosomal DNA, as judged by limited formaldehyde cross-linking of living cells. The physical interaction of kin17 protein with components of the nuclear matrix was confirmed and visualized by indirect immunofluorescence and immunoelectron microscopy. Our results indicate that, during S phase, a fraction of the human kin17 protein preferentially associates with the nuclear matrix, a fundamentally non-chromatin higher order nuclear structure, and to chromatin DNA in the presence of DNA damage.
Figures






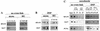
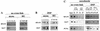
Similar articles
-
The human stress-activated protein kin17 belongs to the multiprotein DNA replication complex and associates in vivo with mammalian replication origins.Mol Cell Biol. 2005 May;25(9):3814-30. doi: 10.1128/MCB.25.9.3814-3830.2005. Mol Cell Biol. 2005. PMID: 15831485 Free PMC article.
-
Participation of kin17 protein in replication factories and in other DNA transactions mediated by high molecular weight nuclear complexes.Mol Cancer Res. 2003 May;1(7):519-31. Mol Cancer Res. 2003. PMID: 12754299
-
Ionizing radiation triggers chromatin-bound kin17 complex formation in human cells.J Biol Chem. 2002 May 24;277(21):19156-65. doi: 10.1074/jbc.M200321200. Epub 2002 Mar 5. J Biol Chem. 2002. PMID: 11880372
-
Nuclear matrix support of DNA replication.J Cell Biochem. 2005 Dec 1;96(5):951-61. doi: 10.1002/jcb.20610. J Cell Biochem. 2005. PMID: 16167334 Review.
-
Nuclear matrix and the cell cycle.Int Rev Cytol. 1995;162B:377-403. doi: 10.1016/s0074-7696(08)62622-4. Int Rev Cytol. 1995. PMID: 8557492 Review.
Cited by
-
Kin17 facilitates multiple double-strand break repair pathways that govern B cell class switching.Sci Rep. 2016 Nov 17;6:37215. doi: 10.1038/srep37215. Sci Rep. 2016. PMID: 27853268 Free PMC article.
-
Roles and regulatory mechanisms of KIN17 in cancers (Review).Oncol Lett. 2023 Feb 17;25(4):137. doi: 10.3892/ol.2023.13723. eCollection 2023 Apr. Oncol Lett. 2023. PMID: 36909374 Free PMC article. Review.
-
Kin17 knockdown suppresses the migration and invasion of cervical cancer cells through NF-κB-Snail pathway.Int J Clin Exp Pathol. 2020 Mar 1;13(3):607-615. eCollection 2020. Int J Clin Exp Pathol. 2020. PMID: 32269702 Free PMC article.
-
Onvansertib and Navitoclax Combination as a New Therapeutic Option for Mucinous Ovarian Carcinoma.Int J Mol Sci. 2025 Jan 8;26(2):472. doi: 10.3390/ijms26020472. Int J Mol Sci. 2025. PMID: 39859203 Free PMC article.
-
A genetic screen in C. elegans reveals roles for KIN17 and PRCC in maintaining 5' splice site identity.PLoS Genet. 2022 Feb 10;18(2):e1010028. doi: 10.1371/journal.pgen.1010028. eCollection 2022 Feb. PLoS Genet. 2022. PMID: 35143478 Free PMC article.
References
-
- Boulikas T. (1996) Common structural features of replication origins in all life forms. J. Cell. Biochem., 60, 297–388. - PubMed
-
- Boulikas T. (1995) Chromatin domains and the prediction of MAR sequences. Int. Rev. Cytol., 162A, 279–388. - PubMed
-
- Cook P.R. (1988) The nucleoskeleton: artefact, passive framework or active site? J. Cell Sci., 90, 1–6. - PubMed
-
- Berezney R., Mortillaro,M.J., Ma,H., Wei,X. and Samarabandu,J. (1995) The nuclear matrix: a structural milieu for genomic function. Int. Rev. Cytol., 162A, 1–65. - PubMed
-
- Nickerson J.A. (2001) Experimental observations of a nuclear matrix. J. Cell Sci., 114, 463–474. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

