Heteromeric MAPPIT: a novel strategy to study modification-dependent protein-protein interactions in mammalian cells
- PMID: 12853652
- PMCID: PMC167658
- DOI: 10.1093/nar/gng075
Heteromeric MAPPIT: a novel strategy to study modification-dependent protein-protein interactions in mammalian cells
Abstract
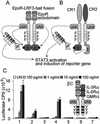
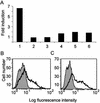
We recently reported a two-hybrid trap for detecting protein-protein interactions in intact mammalian cells (MAPPIT). The bait protein was fused to a STAT recruitment-deficient, homodimeric cytokine receptor and the prey protein to functional STAT recruitment sites. In such a configuration, STAT-dependent responses can be used to monitor a given bait-prey interaction. Using this system, we were able to demonstrate both modification-independent and tyrosine phosphorylation- dependent interactions. Protein modification in this approach is, however, strictly dependent on the receptor-associated JAK tyrosine kinases. We have now extended this concept by using extracellular domains of the heteromeric granulocyte/macrophage colony-stimulating factor receptor (GM-CSFR). Herein, the bait was fused to the (beta)c chain and its modifying enzyme to the GM-CSFRalpha chain (or vice versa). We demonstrate several serine phosphorylation-dependent interactions in the TGFbeta/Smad pathway using the catalytic domains of the ALK4 or ALK6 serine/threonine kinase receptors. In all cases tested, STAT-dependent signaling was completely abolished when mutant baits were used wherein critical serine residues were replaced by alanines. This approach operates both in transient and stable expression systems and may not be limited to serine phosphorylation but has the potential for studying various different types of protein modification-dependent interactions in intact cells.
Figures

References
-
- Fields S. and Song,O. (1989) A novel genetic system to detect protein-protein interactions. Nature, 340, 245–246. - PubMed
-
- Broder Y.C., Katz,S. and Aronheim,A. (1998) The ras recruitment system, a novel approach to the study of protein-protein interactions. Curr. Biol., 8, 1121–1124. - PubMed
-
- Osborne M.A., Dalton,S. and Kochan,J.P. (1995) The yeast tribrid system—genetic detection of trans-phosphorylated ITAM-SH2-interactions. Biotechnology, 13, 1474–1478. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

