The dimer initiation sequence stem-loop of human immunodeficiency virus type 1 is dispensable for viral replication in peripheral blood mononuclear cells
- PMID: 12857902
- PMCID: PMC165254
- DOI: 10.1128/jvi.77.15.8329-8335.2003
The dimer initiation sequence stem-loop of human immunodeficiency virus type 1 is dispensable for viral replication in peripheral blood mononuclear cells
Abstract
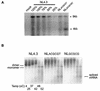
Human immunodeficiency virus type 1 (HIV-1) contains two copies of genomic RNA that are noncovalently linked via a palindrome sequence within the dimer initiation site (DIS) stem-loop. In contrast to the current paradigm that the DIS stem or stem-loop is critical for HIV-1 infectivity, which arose from studies using T-cell lines, we demonstrate here that HIV-1 mutants with deletions in the DIS stem-loop are replication competent in peripheral blood mononuclear cells (PBMCs). The DIS mutants contained either the wild-type (5'GCGCGC3') or an arbitrary (5'ACGCGT3') palindrome sequence in place of the 39-nucleotide DIS stem-loop (NL(CGCGCG) and NL(ACGCGT)). These DIS mutants were replication defective in SupT1 cells, concurring with the current model in which DIS mutants are replication defective in T-cell lines. All of the HIV-1 DIS mutants were replication competent in PBMCs over a 40-day infection period and had retained their respective DIS mutations at 40 days postinfection. Although the stability of the virion RNA dimer was not affected by our DIS mutations, the RNA dimers exhibited a diffuse migration profile when compared to the wild type. No defect in protein processing of the Gag and GagProPol precursor proteins was found in the DIS mutants. Our data provide direct evidence that the DIS stem-loop is dispensable for viral replication in PBMCs and that the requirement of the DIS stem-loop in HIV-1 replication is cell type dependent.
Figures
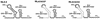
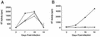
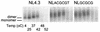

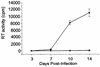
Similar articles
-
Primary T-lymphocytes rescue the replication of HIV-1 DIS RNA mutants in part by facilitating reverse transcription.Nucleic Acids Res. 2008 Mar;36(5):1578-88. doi: 10.1093/nar/gkm1149. Epub 2008 Jan 23. Nucleic Acids Res. 2008. PMID: 18216043 Free PMC article.
-
A short sequence motif in the 5' leader of the HIV-1 genome modulates extended RNA dimer formation and virus replication.J Biol Chem. 2014 Dec 19;289(51):35061-74. doi: 10.1074/jbc.M114.621425. Epub 2014 Nov 3. J Biol Chem. 2014. PMID: 25368321 Free PMC article.
-
Role of the DIS hairpin in replication of human immunodeficiency virus type 1.J Virol. 1996 Oct;70(10):6723-32. doi: 10.1128/JVI.70.10.6723-6732.1996. J Virol. 1996. PMID: 8794309 Free PMC article.
-
A structure-based approach for targeting the HIV-1 genomic RNA dimerization initiation site.Biochimie. 2007 Oct;89(10):1195-203. doi: 10.1016/j.biochi.2007.03.003. Epub 2007 Mar 12. Biochimie. 2007. PMID: 17434658 Review.
-
Methodological approaches to studying phase separation and HIV-1 replication: Current and future perspectives.Methods. 2024 Sep;229:147-155. doi: 10.1016/j.ymeth.2024.07.002. Epub 2024 Jul 15. Methods. 2024. PMID: 39002735 Review.
Cited by
-
The A-rich RNA sequences of HIV-1 pol are important for the synthesis of viral cDNA.Nucleic Acids Res. 2009 Feb;37(3):945-56. doi: 10.1093/nar/gkn1015. Epub 2008 Dec 23. Nucleic Acids Res. 2009. PMID: 19106143 Free PMC article.
-
X4 and R5 HIV-1 have distinct post-entry requirements for uracil DNA glycosylase during infection of primary cells.J Biol Chem. 2010 Jun 11;285(24):18603-14. doi: 10.1074/jbc.M109.090126. Epub 2010 Apr 6. J Biol Chem. 2010. PMID: 20371602 Free PMC article.
-
Delineation of the preferences and requirements of the human immunodeficiency virus type 1 dimerization initiation signal by using an in vivo cell-based selection approach.J Virol. 2010 Jul;84(13):6866-75. doi: 10.1128/JVI.01930-09. Epub 2010 Apr 21. J Virol. 2010. PMID: 20410279 Free PMC article.
-
Effects of a single amino acid substitution within the p2 region of human immunodeficiency virus type 1 on packaging of spliced viral RNA.J Virol. 2003 Dec;77(24):12986-95. doi: 10.1128/jvi.77.24.12986-12995.2003. J Virol. 2003. PMID: 14645555 Free PMC article.
-
Sequence analysis of the dimerization initiation site of concordant and discordant viral variants superinfecting HIV type 1 patients.AIDS Res Hum Retroviruses. 2011 Nov;27(11):1231-5. doi: 10.1089/AID.2011.0010. Epub 2011 May 25. AIDS Res Hum Retroviruses. 2011. PMID: 21453132 Free PMC article.
References
-
- Berkhout, B. 1996. Structure and function of the human immunodeficiency virus leader RNA. Prog. Nucleic Acid Res. Mol. Biol. 54:1-34. - PubMed
-
- Berkhout, B., M. Ooms, N. Beerens, H. Huthoff, E. Southern, and K. Verhoef. 2002. In vitro evidence that the untranslated leader of the HIV-1 genome is an RNA checkpoint that regulates multiple functions through conformational changes. J. Biol. Chem. 277:19967-19975. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

