Use of RNA interference and complementation to study the function of the Drosophila and human 26S proteasome subunit S13
- PMID: 12861018
- PMCID: PMC165711
- DOI: 10.1128/MCB.23.15.5320-5330.2003
Use of RNA interference and complementation to study the function of the Drosophila and human 26S proteasome subunit S13
Abstract
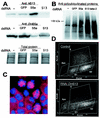
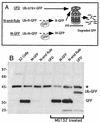
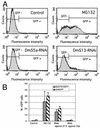
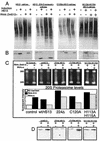
The S13 subunit (also called Pad1, Rpn11, and MPR1) is a component of the 19S complex, a regulatory complex essential for the ubiquitin-dependent proteolytic activity of the 26S proteasome. To address the functional role of S13, we combined double-stranded RNA interference (RNAi) against the Drosophila proteasome subunit DmS13 with expression of wild-type and mutant forms of the homologous human gene, HS13. These studies show that DmS13 is essential for 26S function. Loss of the S13 subunit in metazoan cells leads to increased levels of ubiquitin conjugates, cell cycle defects, DNA overreplication, and apoptosis. In vivo assays using short-lived proteasome substrates confirmed that the 26S ubiquitin-dependent degradation pathway is compromised in S13-depleted cells. In complementation experiments using Drosophila cell lines expressing HS13, wild-type HS13 was found to fully rescue the knockdown phenotype after DmS13 RNAi treatment, while an HS13 containing mutations (H113A-H115A) in the proposed isopeptidase active site was unable to rescue. A mutation within the conserved MPN/JAMM domain (C120A) abolished the ability of HS13 to rescue the Drosophila cells from apoptosis or DNA overreplication. However, the C120A mutant was found to partially restore normal levels of ubiquitin conjugates. The S13 subunit may possess multiple functions, including a deubiquitinylating activity and distinct activities essential for cell cycle progression that require the conserved C120 residue.
Figures


Similar articles
-
Participation of the proteasomal lid subunit Rpn11 in mitochondrial morphology and function is mapped to a distinct C-terminal domain.Biochem J. 2004 Jul 1;381(Pt 1):275-85. doi: 10.1042/BJ20040008. Biochem J. 2004. PMID: 15018611 Free PMC article.
-
Role of Rpn11 metalloprotease in deubiquitination and degradation by the 26S proteasome.Science. 2002 Oct 18;298(5593):611-5. doi: 10.1126/science.1075898. Epub 2002 Aug 15. Science. 2002. PMID: 12183636
-
Deletion of proteasomal subunit S5a/Rpn10/p54 causes lethality, multiple mitotic defects and overexpression of proteasomal genes in Drosophila melanogaster.J Cell Sci. 2003 Mar 15;116(Pt 6):1023-33. doi: 10.1242/jcs.00332. J Cell Sci. 2003. PMID: 12584246
-
GFP-labelling of 26S proteasomes in living yeast: insight into proteasomal functions at the nuclear envelope/rough ER.Mol Biol Rep. 1999 Apr;26(1-2):131-5. doi: 10.1023/a:1006973803960. Mol Biol Rep. 1999. PMID: 10363659 Review.
-
Proteins interacting with the 26S proteasome.Cell Mol Life Sci. 2004 Jul;61(13):1589-95. doi: 10.1007/s00018-004-4132-x. Cell Mol Life Sci. 2004. PMID: 15224183 Free PMC article. Review.
Cited by
-
Disruption of Rpn4-induced proteasome expression in Saccharomyces cerevisiae reduces cell viability under stressed conditions.Genetics. 2008 Dec;180(4):1945-53. doi: 10.1534/genetics.108.094524. Epub 2008 Oct 1. Genetics. 2008. PMID: 18832351 Free PMC article.
-
Adaptability of the ubiquitin-proteasome system to proteolytic and folding stressors.J Cell Biol. 2021 Mar 1;220(3):e201912041. doi: 10.1083/jcb.201912041. J Cell Biol. 2021. PMID: 33382395 Free PMC article.
-
Deubiquitination Reactions on the Proteasome for Proteasome Versatility.Int J Mol Sci. 2020 Jul 27;21(15):5312. doi: 10.3390/ijms21155312. Int J Mol Sci. 2020. PMID: 32726943 Free PMC article. Review.
-
Hsp83 loss suppresses proteasomal activity resulting in an upregulation of caspase-dependent compensatory autophagy.Autophagy. 2017 Sep 2;13(9):1573-1589. doi: 10.1080/15548627.2017.1339004. Epub 2017 Aug 14. Autophagy. 2017. PMID: 28806103 Free PMC article.
-
A proteasome for all occasions.FEBS Lett. 2007 Jun 19;581(15):2854-61. doi: 10.1016/j.febslet.2007.03.053. Epub 2007 Mar 30. FEBS Lett. 2007. PMID: 17418826 Free PMC article. Review.
References
-
- Arioka, M., M. Kouhashi, K. Yoda, A. Takatsuki, M. Yamasaki, and K. Kitamoto. 1998. Multidrug resistance phenotype conferred by overexpressing bfr2+/pad1+/sks1+ or pap1+ genes and mediated by bfr1+ gene product, a structural and functional homologue of P-glycoprotein in Schizosaccharomyces pombe. Biosci. Biotechnol. Biochem. 62:390-392. - PubMed
-
- Bosher, J. M., and M. Labouesse. 2000. RNA interference: genetic wand and genetic watchdog. Nat. Cell Biol. 2:E31-E36. - PubMed
-
- Braun, B. C., M. Glickman, R. Kraft, B. Dahlmann, P. M. Kloetzel, D. Finley, and M. Schmidt. 1999. The base of the proteasome regulatory particle exhibits chaperone-like activity. Nat. Cell Biol. 1:221-226. - PubMed
-
- Cope, G. A., G. S. Suh, L. Aravind, S. E. Schwarz, S. L. Zipursky, E. V. Koonin, and R. J. Deshaies. 2002. Role of predicted metalloprotease motif of Jab1/Csn5 in cleavage of Nedd8 from Cul1. Science 298:608-611. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
