Cytoplasmically retargeted HSV1-tk/GFP reporter gene mutants for optimization of noninvasive molecular-genetic imaging
- PMID: 12869307
- PMCID: PMC1502405
- DOI: 10.1016/S1476-5586(03)80056-8
Cytoplasmically retargeted HSV1-tk/GFP reporter gene mutants for optimization of noninvasive molecular-genetic imaging
Abstract
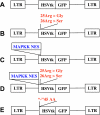
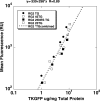
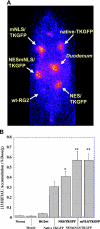
To optimize the sensitivity of imaging HSV1-tk/GFP reporter gene expression, a series of HSV1-tk/GFP mutants was developed with altered nuclear localization and better cellular enzymatic activity, compared to that of the native HSV1-tk/GFP fusion protein (HSV1-tk/GFP). Several modifications of HSV1-tk/GFP reporter gene were performed, including targeted inactivating mutations in the nuclear localization signal (NLS), the addition of a nuclear export signal (NES), a combination of both mutation types, and a truncation of the first 135 bp of the native hsv1-tk coding sequence containing a "cryptic" testicular promoter and the NLS. A recombinant HSV1-tk/GFP protein and a highly sensitive sandwich enzyme-linked immunosorbent assay for HSV1-tk/GFP were developed to quantitate the amount of reporter gene product in different assays to allow normalization of the data. These different mutations resulted in various degrees of nuclear clearance, predominant cytoplasmic distribution, and increased total cellular enzymatic activity of the HSV1-tk/GFP mutants, compared to native HSV1-tk/GFP when expressed at the same levels. This appears to be the result of improved metabolic bioavailability of cytoplasmically retargeted mutant HSV1-tk/GFP enzymes for reaction with the radiolabeled probe (e.g., FIAU). The analysis of enzymatic properties of different HSV1-tk/GFP mutants using FIAU as a substrate revealed no significant differences from that of the native HSV1-tk/GFP. Improved total cellular enzymatic activity of cytoplasmically retargeted HSV1-tk/GFP mutants observed in vitro was confirmed by noninvasive imaging of transduced subcutaneous tumor xenografts bearing these reporters using [(131)I]FIAU and a gamma-camera.
Figures



Similar articles
-
A novel triple-modality reporter gene for whole-body fluorescent, bioluminescent, and nuclear noninvasive imaging.Eur J Nucl Med Mol Imaging. 2004 May;31(5):740-51. doi: 10.1007/s00259-003-1441-5. Epub 2004 Mar 11. Eur J Nucl Med Mol Imaging. 2004. PMID: 15014901
-
Herpes simplex virus thymidine kinase as a marker/reporter gene for PET imaging of gene therapy.Q J Nucl Med. 1999 Jun;43(2):163-9. Q J Nucl Med. 1999. PMID: 10429512 Review.
-
PET of cardiac transgene expression: comparison of 2 approaches based on herpesviral thymidine kinase reporter gene.J Nucl Med. 2004 Nov;45(11):1917-23. J Nucl Med. 2004. PMID: 15534063
-
Myocardial kinetics of reporter probe 124I-FIAU in isolated perfused rat hearts after in vivo adenoviral transfer of herpes simplex virus type 1 thymidine kinase reporter gene.J Nucl Med. 2005 Jan;46(1):98-105. J Nucl Med. 2005. PMID: 15632039
-
HSV1-TK/GFP/Fluc.2008 Oct 21 [updated 2008 Dec 1]. In: Molecular Imaging and Contrast Agent Database (MICAD) [Internet]. Bethesda (MD): National Center for Biotechnology Information (US); 2004–2013. 2008 Oct 21 [updated 2008 Dec 1]. In: Molecular Imaging and Contrast Agent Database (MICAD) [Internet]. Bethesda (MD): National Center for Biotechnology Information (US); 2004–2013. PMID: 20641334 Free Books & Documents. Review.
Cited by
-
PET imaging of HSV1-tk mutants with acquired specificity toward pyrimidine- and acycloguanosine-based radiotracers.Eur J Nucl Med Mol Imaging. 2009 Aug;36(8):1273-82. doi: 10.1007/s00259-009-1089-x. Epub 2009 Mar 4. Eur J Nucl Med Mol Imaging. 2009. PMID: 19259663 Free PMC article.
-
Human breast tumor cells express multimodal imaging reporter genes.Mol Imaging Biol. 2008 Sep;10(5):253-63. doi: 10.1007/s11307-008-0147-2. Epub 2008 Jun 17. Mol Imaging Biol. 2008. PMID: 18560942
-
ERCC2 Helicase Domain Mutations Confer Nucleotide Excision Repair Deficiency and Drive Cisplatin Sensitivity in Muscle-Invasive Bladder Cancer.Clin Cancer Res. 2019 Feb 1;25(3):977-988. doi: 10.1158/1078-0432.CCR-18-1001. Epub 2018 Jul 6. Clin Cancer Res. 2019. PMID: 29980530 Free PMC article.
-
Monitoring the efficacy of adoptively transferred prostate cancer-targeted human T lymphocytes with PET and bioluminescence imaging.J Nucl Med. 2008 Jul;49(7):1162-70. doi: 10.2967/jnumed.107.047324. Epub 2008 Jun 13. J Nucl Med. 2008. PMID: 18552144 Free PMC article.
-
Imaging of Sleeping Beauty-Modified CD19-Specific T Cells Expressing HSV1-Thymidine Kinase by Positron Emission Tomography.Mol Imaging Biol. 2016 Dec;18(6):838-848. doi: 10.1007/s11307-016-0971-8. Mol Imaging Biol. 2016. PMID: 27246312 Free PMC article.
References
-
- Bengel FM, Anton M, Avril N, Brill T, Nguyen N, Haubner R, Gleiter E, Gansbacher B, Schwaiger M. Uptake of radiolabeled 2′-fluoro-2′-deoxy-5-iodo-1-beta-D-arabinofuranosyluracil in cardiac cells after adenoviral transfer of the herpesvirus thymidine kinase gene: the cellular basis for cardiac gene imaging. Circulation. 2000;102:948–950. - PubMed
-
- Black ME, Kokoris MS, Sabo P. Herpes simplex virus-1 thymidine kinase mutants created by semi-random sequence mutagenesis improve prodrug-mediated tumor cell killing. Cancer Res. 2001;61:3022–3026. - PubMed
-
- Black ME, Rechtin TM, Drake RR. Effect on substrate binding of an alteration at the conserved aspartic acid-162 in herpes simplex virus type 1 thymidine kinase. J Gen Virol. 1996;77(Pt 7):1521–1527. - PubMed
-
- Brown DG, Visse R, Sandhu G, Davies A, Rizkallah P, Melitz C, Summers W, Sanderson MR. Crystal structures of the thymidine kinase from herpes simplex virus type-1 in complex with deoxythymidine and ganciclovir. Nat Struct Biol. 1995;2:876–881. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

