Use of nucleotide analogs by class I and class II CCA-adding enzymes (tRNA nucleotidyltransferase): deciphering the basis for nucleotide selection
- PMID: 12869708
- PMCID: PMC1370463
- DOI: 10.1261/rna.2110903
Use of nucleotide analogs by class I and class II CCA-adding enzymes (tRNA nucleotidyltransferase): deciphering the basis for nucleotide selection
Abstract
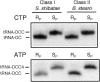
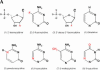
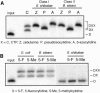
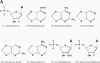
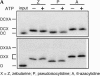
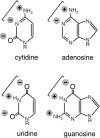
We explored the specificity and nature of the nucleotide-binding pocket of the CCA-adding enzyme (tRNA nucleotidyltransferase) by using CTP and ATP analogs as substrates for a panel of class I and class II enzymes. Overall, class I and class II enzymes displayed remarkably similar substrate requirements, implying that the mechanism of CCA addition is conserved between enzyme classes despite the absence of obvious sequence homology outside the active site signature sequence. CTP substrates are more tolerant of base modifications than ATP substrates, but sugar modifications prevent incorporation of both CTP and ATP analogs by class I and class II enzymes. Use of CTP analogs (zebularine, pseudoisocytidine, 6-azacytidine, but not 6-azauridine) suggests that base modifications generally do not interfere with recognition or incorporation of CTP analogs by either class I or class II enzymes, and that UTP is excluded because N-3 is a positive determinant and/or O-4 is an antideterminant. Use of ATP analogs (N6-methyladenosine, diaminopurine, purine, 2-aminopurine, and 7-deaza-adenosine, but not guanosine, deoxyadenosine, 2'-O-methyladenosine, 2'-deoxy-2'-fluoroadenosine, or inosine) suggests that base modifications generally do not interfere with recognition or incorporation of ATP analogs by either class I or class II enzymes, and that GTP is excluded because N-1 is a positive determinant and/or the 2-amino and 6-keto groups are antideterminants. We also found that the 3'-terminal sequence of the growing tRNA substrate can affect the efficiency or specificity of subsequent nucleotide addition. Our data set should allow rigorous evaluation of structural hypotheses for nucleotide selection based on existing and future crystal structures.
Figures










Similar articles
-
Poly(C) synthesis by class I and class II CCA-adding enzymes.Biochemistry. 2002 Apr 9;41(14):4521-32. doi: 10.1021/bi0120953. Biochemistry. 2002. PMID: 11926813
-
CCA addition by tRNA nucleotidyltransferase: polymerization without translocation?EMBO J. 1998 Jun 1;17(11):3197-206. doi: 10.1093/emboj/17.11.3197. EMBO J. 1998. PMID: 9606201 Free PMC article.
-
Divergent evolutions of trinucleotide polymerization revealed by an archaeal CCA-adding enzyme structure.EMBO J. 2003 Nov 3;22(21):5918-27. doi: 10.1093/emboj/cdg563. EMBO J. 2003. PMID: 14592988 Free PMC article.
-
Molecular mechanisms of template-independent RNA polymerization by tRNA nucleotidyltransferases.Front Genet. 2014 Feb 17;5:36. doi: 10.3389/fgene.2014.00036. eCollection 2014. Front Genet. 2014. PMID: 24596576 Free PMC article. Review.
-
A story with a good ending: tRNA 3'-end maturation by CCA-adding enzymes.Curr Opin Struct Biol. 2006 Feb;16(1):12-7. doi: 10.1016/j.sbi.2005.12.001. Epub 2005 Dec 20. Curr Opin Struct Biol. 2006. PMID: 16364630 Review.
Cited by
-
[3'-32P]-labeling tRNA with nucleotidyltransferase for assaying aminoacylation and peptide bond formation.Methods. 2008 Feb;44(2):74-80. doi: 10.1016/j.ymeth.2007.08.001. Methods. 2008. PMID: 18241789 Free PMC article.
-
Recognition of guanosine by dissimilar tRNA methyltransferases.RNA. 2012 Sep;18(9):1687-701. doi: 10.1261/rna.032029.111. Epub 2012 Jul 30. RNA. 2012. PMID: 22847817 Free PMC article.
-
Reengineering CCA-adding enzymes to function as (U,G)- or dCdCdA-adding enzymes or poly(C,A) and poly(U,G) polymerases.Proc Natl Acad Sci U S A. 2007 Jan 2;104(1):54-9. doi: 10.1073/pnas.0606961104. Epub 2006 Dec 19. Proc Natl Acad Sci U S A. 2007. PMID: 17179213 Free PMC article.
-
Peptide release on the ribosome depends critically on the 2' OH of the peptidyl-tRNA substrate.RNA. 2008 Aug;14(8):1526-31. doi: 10.1261/rna.1057908. Epub 2008 Jun 20. RNA. 2008. PMID: 18567817 Free PMC article.
-
On the role of a conserved, potentially helix-breaking residue in the tRNA-binding alpha-helix of archaeal CCA-adding enzymes.RNA. 2008 Jul;14(7):1284-9. doi: 10.1261/rna.1060308. Epub 2008 May 21. RNA. 2008. PMID: 18495940 Free PMC article.
References
-
- Abramovitz, D.L., Friedman, R.A., and Pyle, A.M. 1996. Catalytic role of 2′-hydroxyl groups within a group II intron active site. Science 271: 1410–1413. - PubMed
-
- Aebi, M., Kirchner, G., Chen, J.Y., Vijayraghavan, U., Jacobson, A., Martin, N.C., and Abelson, J. 1990. Isolation of a temperature-sensitive mutant with an altered tRNA nucleotidyltransferase and cloning of the gene encoding tRNA nucleotidyltransferase in the yeast Saccharomyces cerevisiae. J. Biol. Chem. 265: 16216–16220. - PubMed
-
- Cho, H.D., Tomita, K., Suzuki, T., and Weiner, A.M. 2002. U2 small nuclear RNA is a substrate for the CCA-adding enzyme (tRNA nucleotidyltransferase). J. Biol. Chem. 277: 3447–3455. - PubMed
-
- Cramer, F., Faulhammer, H., von der Haar, F., Sprinzl, M., and Sternbach, H. 1975. Aminoacyl-tRNA synthetases from baker’s yeast: Reacting site of enzymatic aminoacylation is not uniform for all tRNAs. FEBS Lett. 56: 212–214. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
