PorA represents the major cell wall channel of the Gram-positive bacterium Corynebacterium glutamicum
- PMID: 12896997
- PMCID: PMC166458
- DOI: 10.1128/JB.185.16.4779-4786.2003
PorA represents the major cell wall channel of the Gram-positive bacterium Corynebacterium glutamicum
Abstract
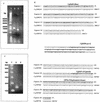
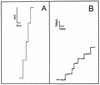
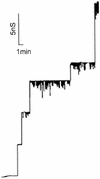
The cell wall of the gram-positive bacterium Corynebacterium glutamicum contains a channel (porin) for the passage of hydrophilic solutes. The channel-forming polypeptide PorA is a 45-amino-acid acidic polypeptide with an excess of four negatively charged amino acids, which is encoded by the 138-bp gene porA. porA was deleted from the chromosome of C.glutamicum wild-type strain ATCC 13032 to obtain mutant ATCC 13032deltaporA. Southern blot analysis demonstrated that porA was deleted. Lipid bilayer experiments revealed that PorA was not present in the cell wall of the mutant strain. Searches within the known chromosome of C. glutamicum by using National Center for Biotechnology Information BLAST and reverse transcription-PCR showed that no other PorA-like protein is encoded on the chromosome or is expressed in the deletion strain. The porA deletion strain exhibited slower growth and longer growth times than the C. glutamicum wild-type strain. Experiments with different antibiotics revealed that the susceptibility of the mutant strain was much lower than that of the wild-type C. glutamicum strain. The results presented here suggest that PorA represents a major hydrophilic pathway through the cell wall and that C. glutamicum contains cell wall channels which are not related to PorA.
Figures


Similar articles
-
Identification of an anion-specific channel in the cell wall of the Gram-positive bacterium Corynebacterium glutamicum.Mol Microbiol. 2003 Nov;50(4):1295-308. doi: 10.1046/j.1365-2958.2003.03754.x. Mol Microbiol. 2003. PMID: 14622416
-
Reconstitution experiments and gene deletions reveal the existence of two-component major cell wall channels in the genus Corynebacterium.J Bacteriol. 2010 Feb;192(3):786-800. doi: 10.1128/JB.01142-09. Epub 2009 Dec 4. J Bacteriol. 2010. PMID: 19966008 Free PMC article.
-
Biochemical and biophysical characterization of the cell wall porin of Corynebacterium glutamicum: the channel is formed by a low molecular mass polypeptide.Biochemistry. 1998 Oct 27;37(43):15024-32. doi: 10.1021/bi980961e. Biochemistry. 1998. PMID: 9790664
-
Mycomembrane and S-layer: two important structures of Corynebacterium glutamicum cell envelope with promising biotechnology applications.J Biotechnol. 2003 Sep 4;104(1-3):55-67. doi: 10.1016/s0168-1656(03)00163-9. J Biotechnol. 2003. PMID: 12948629 Review.
-
Promoters of Corynebacterium glutamicum.J Biotechnol. 2003 Sep 4;104(1-3):311-23. doi: 10.1016/s0168-1656(03)00155-x. J Biotechnol. 2003. PMID: 12948648 Review.
Cited by
-
Corynebacterium diphtheriae: identification and characterization of a channel-forming protein in the cell wall.J Bacteriol. 2007 Nov;189(21):7709-19. doi: 10.1128/JB.00864-07. Epub 2007 Aug 24. J Bacteriol. 2007. PMID: 17720794 Free PMC article.
-
Click-chemistry approach to study mycoloylated proteins: Evidence for PorB and PorC porins mycoloylation in Corynebacterium glutamicum.PLoS One. 2017 Feb 15;12(2):e0171955. doi: 10.1371/journal.pone.0171955. eCollection 2017. PLoS One. 2017. PMID: 28199365 Free PMC article.
-
Corynebacterium jeikeium jk0268 constitutes for the 40 amino acid long PorACj, which forms a homooligomeric and anion-selective cell wall channel.PLoS One. 2013 Oct 8;8(10):e75651. doi: 10.1371/journal.pone.0075651. eCollection 2013. PLoS One. 2013. PMID: 24116064 Free PMC article.
-
Identification and characterization of smallest pore-forming protein in the cell wall of pathogenic Corynebacterium urealyticum DSM 7109.BMC Biochem. 2018 May 9;19(1):3. doi: 10.1186/s12858-018-0093-9. BMC Biochem. 2018. PMID: 29743008 Free PMC article.
-
Uptake pathways of anionic and cationic photosensitizers into bacteria.Photochem Photobiol Sci. 2009 Jun;8(6):788-95. doi: 10.1039/b809624d. Epub 2009 Mar 31. Photochem Photobiol Sci. 2009. PMID: 19492106 Free PMC article.
References
-
- Achouak, W., T. Heulin, and J. M. Pages. 2001. Multiple facets of bacterial porins. FEMS Microbiol. Lett. 199:1-7. - PubMed
-
- Barksdale L. 1981. The genus Corynebacterium, p. 1827-1837. In M. P. Starr, H. Stoll, H. G. Truper, A. Balows, and H. G. Schlegel (ed.), The prokaryotes. Springer-Verlag, Berlin, Germany.
-
- Beckers, G., L. Nolden, and A. Burkovski. 2000. Glutamate synthase of Corynebacterium glutamicum is not essential for glutamate synthesis and is regulated by the nitrogen status. Microbiology 147:2961-2970. - PubMed
-
- Benz, R. 2001. Porins—structure and function, p. 227-246. In G. Winkelmann (ed.), Microbial transport systems. WILEY-VCH, Weinheim, Germany.
-
- Benz, R., K. Janko, W. Boos, and P. Lauger. 1978. Formation of large, ion-permeable membrane channels by the matrix protein (porin) of Escherichia coli. Biochim. Biophys. Acta 511:305-319. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

