Molecular analysis of Phr peptide processing in Bacillus subtilis
- PMID: 12897006
- PMCID: PMC166482
- DOI: 10.1128/JB.185.16.4861-4871.2003
Molecular analysis of Phr peptide processing in Bacillus subtilis
Abstract
In Bacillus subtilis, an export-import pathway regulates production of the Phr pentapeptide inhibitors of Rap proteins. Processing of the Phr precursor proteins into the active pentapeptide form is a key event in the initiation of sporulation and competence development. The PhrA (ARNQT) and PhrE (SRNVT) peptides inhibit the RapA and RapE phosphatases, respectively, whose activity is directed toward the Spo0F approximately P intermediate response regulator of the sporulation phosphorelay. The PhrC (ERGMT) peptide inhibits the RapC protein acting on the ComA response regulator for competence with regard to DNA transformation. The structural organization of PhrA, PhrE, and PhrC suggested a role for type I signal peptidases in the processing of the Phr preinhibitor, encoded by the phr genes, into the proinhibitor form. The proinhibitor was then postulated to be cleaved to the active pentapeptide inhibitor by an additional enzyme. In this report, we provide evidence that Phr preinhibitor proteins are subject to only one processing event at the peptide bond on the amino-terminal end of the pentapeptide. This processing event is most likely independent of type I signal peptidase activity. In vivo and in vitro analyses indicate that none of the five signal peptidases of B. subtilis (SipS, SipT, SipU, SipV, and SipW) are indispensable for Phr processing. However, we show that SipV and SipT have a previously undescribed role in sporulation, competence, and cell growth.
Figures




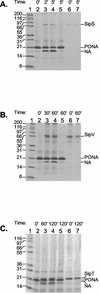

References
-
- Bligh, E. G., and W. J. Dyer. 1959. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37:911-917. - PubMed
-
- Burkholder, W. F., I. Kurtser, and A. D. Grossman. 2001. Replication initiation proteins regulate a developmental checkpoint in Bacillus subtilis. Cell 104:269-279. - PubMed
-
- Carlos, J. L., M. Paetzel, G. Brubaker, A. Karla, C. M. Ashwell, M. O. Lively, G. Cao, P. Bullinger, and R. E. Dalbey. 2000. The role of the membrane-spanning domain of type I signal peptidases in substrate cleavage site selection. J. Biol. Chem. 275:38813-38822. - PubMed
-
- Chatterjee, S., D. Suciu, R. E. Dalbey, P. C. Kahn, and M. Inouye. 1995. Determination of Km and kcat for signal peptidase I using a full length secretory precursor, pro-OmpA-nuclease A. J. Mol. Biol. 245:311-314. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

