Xeroderma pigmentosum group A protein loads as a separate factor onto DNA lesions
- PMID: 12897146
- PMCID: PMC166334
- DOI: 10.1128/MCB.23.16.5755-5767.2003
Xeroderma pigmentosum group A protein loads as a separate factor onto DNA lesions
Abstract
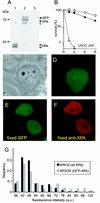
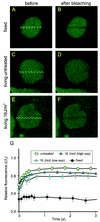
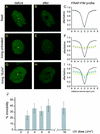
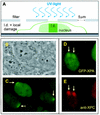
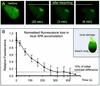
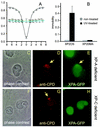
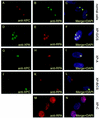
Nucleotide excision repair (NER) is the main DNA repair pathway in mammals for removal of UV-induced lesions. NER involves the concerted action of more than 25 polypeptides in a coordinated fashion. The xeroderma pigmentosum group A protein (XPA) has been suggested to function as a central organizer and damage verifier in NER. How XPA reaches DNA lesions and how the protein is distributed in time and space in living cells are unknown. Here we studied XPA in vivo by using a cell line stably expressing physiological levels of functional XPA fused to green fluorescent protein and by applying quantitative fluorescence microscopy. The majority of XPA moves rapidly through the nucleoplasm with a diffusion rate different from those of other NER factors tested, arguing against a preassembled XPA-containing NER complex. DNA damage induced a transient ( approximately 5-min) immobilization of maximally 30% of XPA. Immobilization depends on XPC, indicating that XPA is not the initial lesion recognition protein in vivo. Moreover, loading of replication protein A on NER lesions was not dependent on XPA. Thus, XPA participates in NER by incorporation of free diffusing molecules in XPC-dependent NER-DNA complexes. This study supports a model for a rapid consecutive assembly of free NER factors, and a relatively slow simultaneous disassembly, after repair.
Figures

Similar articles
-
A truncated human xeroderma pigmentosum complementation group A protein expressed from an adenovirus sensitizes human tumor cells to ultraviolet light and cisplatin.Cancer Res. 2001 Jan 15;61(2):764-70. Cancer Res. 2001. PMID: 11212280
-
Phosphorylation of nucleotide excision repair factor xeroderma pigmentosum group A by ataxia telangiectasia mutated and Rad3-related-dependent checkpoint pathway promotes cell survival in response to UV irradiation.Cancer Res. 2006 Mar 15;66(6):2997-3005. doi: 10.1158/0008-5472.CAN-05-3403. Cancer Res. 2006. PMID: 16540648 Free PMC article.
-
Cell-type-specific consequences of nucleotide excision repair deficiencies: Embryonic stem cells versus fibroblasts.DNA Repair (Amst). 2008 Oct 1;7(10):1659-69. doi: 10.1016/j.dnarep.2008.06.009. Epub 2008 Jul 26. DNA Repair (Amst). 2008. PMID: 18634906
-
Xeroderma Pigmentosa Group A (XPA), Nucleotide Excision Repair and Regulation by ATR in Response to Ultraviolet Irradiation.Adv Exp Med Biol. 2017;996:41-54. doi: 10.1007/978-3-319-56017-5_4. Adv Exp Med Biol. 2017. PMID: 29124689 Free PMC article. Review.
-
Mechanism and regulation of DNA damage recognition in mammalian nucleotide excision repair.Enzymes. 2019;45:99-138. doi: 10.1016/bs.enz.2019.06.004. Epub 2019 Jul 8. Enzymes. 2019. PMID: 31627884 Review.
Cited by
-
Sequential recruitment of the repair factors during NER: the role of XPG in initiating the resynthesis step.EMBO J. 2008 Jan 9;27(1):155-67. doi: 10.1038/sj.emboj.7601948. Epub 2007 Dec 13. EMBO J. 2008. PMID: 18079701 Free PMC article.
-
Other proteins interacting with XP proteins.Adv Exp Med Biol. 2008;637:103-12. doi: 10.1007/978-0-387-09599-8_11. Adv Exp Med Biol. 2008. PMID: 19181115 Free PMC article. Review. No abstract available.
-
DDB2 promotes chromatin decondensation at UV-induced DNA damage.J Cell Biol. 2012 Apr 16;197(2):267-81. doi: 10.1083/jcb.201106074. Epub 2012 Apr 9. J Cell Biol. 2012. PMID: 22492724 Free PMC article.
-
DNA damage response.Cold Spring Harb Perspect Biol. 2011 Jan 1;3(1):a000745. doi: 10.1101/cshperspect.a000745. Cold Spring Harb Perspect Biol. 2011. PMID: 20980439 Free PMC article. Review.
-
Mechanism of action of nucleotide excision repair machinery.Biochem Soc Trans. 2022 Feb 28;50(1):375-386. doi: 10.1042/BST20210246. Biochem Soc Trans. 2022. PMID: 35076656 Free PMC article. Review.
References
-
- Aboussekhra, A., M. Biggerstaff, M. K. K. Shivji, J. A. Vilpo, V. Moncollin, V. N. Podust, M. Protic, U. Hubscher, J.-M. Egly, and R. D. Wood. 1995. Mammalian DNA nucleotide excision repair reconstituted with purified components. Cell 80:859-868. - PubMed
-
- Araki, M., C. Masutani, M. Takemura, A. Uchida, K. Sugasawa, J. Kondoh, Y. Ohkuma, and F. Hanaoka. 2001. Centrosome protein centrin 2/caltractin 1 is part of the xeroderma pigmentosum group C complex that initiates global genome nucleotide excision repair. J. Biol. Chem. 276:18665-18672. - PubMed
-
- Araujo, S. J., F. Tirode, F. Coin, H. Pospiech, J. E. Syvaoja, M. Stucki, U. Hubscher, J. M. Egly, and R. D. Wood. 2000. Nucleotide excision repair of DNA with recombinant human proteins: definition of the minimal set of factors, active forms of TFIIH, and modulation by CAK. Genes Dev. 14:349-359. - PMC - PubMed
-
- Araujo, S. J., and R. D. Wood. 1999. Protein complexes in nucleotide excision repair. Mutat. Res. 435:23-33. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
