Ribozyme-mediated revision of RNA and DNA
- PMID: 12897196
- PMCID: PMC166303
- DOI: 10.1172/JCI19386
Ribozyme-mediated revision of RNA and DNA
Figures
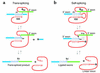

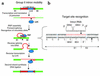

References
-
- Cech, T.R. 1993. Structure and mechanism of the large catalytic RNAs: Group I and group II introns and ribonuclease P. In The RNA World. R.F. Gesteland and J.F. Atkins, editors. Cold Spring Harbor Laboratory Press. Cold Spring Harbor, New York, USA. 239–269.
-
- Sullenger BA, Cech TR. Ribozyme-mediated repair of defective mRNA by targeted trans-splicing. Nature. 1994;371:619–622. - PubMed
-
- Puttaraju M, Jamison SF, Mansfield SG, Garcia-Blanco MA, Mitchell LG. Spliceosome-mediated RNA trans-splicing as a tool for gene therapy. Nat. Biotech. 1999;17:246–252. - PubMed
-
- Guo H, et al. Group II introns designed to insert into therapeutically relevant DNA target sites in human cells. Science. 2000;289:452–457. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

