Microbial ecology of an extreme acidic environment, the Tinto River
- PMID: 12902280
- PMCID: PMC169134
- DOI: 10.1128/AEM.69.8.4853-4865.2003
Microbial ecology of an extreme acidic environment, the Tinto River
Erratum in
- Appl Environ Microbiol. 2003 Nov;69(11):6959
Abstract
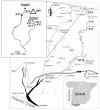
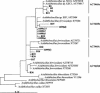
The Tinto River (Huelva, southwestern Spain) is an extreme environment with a rather constant acidic pH along the entire river and a high concentration of heavy metals. The extreme conditions of the Tinto ecosystem are generated by the metabolic activity of chemolithotrophic microorganisms thriving in the rich complex sulfides of the Iberian Pyrite Belt. Molecular ecology techniques were used to analyze the diversity of this microbial community. The community's composition was studied by denaturing gradient gel electrophoresis (DGGE) using 16S rRNA and by 16S rRNA gene amplification. A good correlation between the two approaches was found. Comparative sequence analysis of DGGE bands showed the presence of organisms related to Leptospirillum spp., Acidithiobacillus ferrooxidans, Acidiphilium spp., "Ferrimicrobium acidiphilum," Ferroplasma acidiphilum, and Thermoplasma acidophilum. The different phylogenetic groups were quantified by fluorescent in situ hybridization with a set of rRNA-targeted oligonucleotide probes. More than 80% of the cells were affiliated with the domain Bacteria, with only a minor fraction corresponding to Archaea. Members of Leptospirillum ferrooxidans, Acidithiobacillus ferrooxidans, and Acidiphilium spp., all related to the iron cycle, accounted for most of the prokaryotic microorganisms detected. Different isolates of these microorganisms were obtained from the Tinto ecosystem, and their physiological properties were determined. Given the physicochemical characteristics of the habitat and the physiological properties and relative concentrations of the different prokaryotes found in the river, a model for the Tinto ecosystem based on the iron cycle is suggested.
Figures




References
-
- Amaral-Zettler, L. A., F. Gómez, E. Zettler, B. G. Keenan, R. Amils, and M. L. Sogin. 2002. Eukaryotic diversity in Spain's River of Fire. Nature 417:137. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

