Is the in situ accessibility of the 16S rRNA of Escherichia coli for Cy3-labeled oligonucleotide probes predicted by a three-dimensional structure model of the 30S ribosomal subunit?
- PMID: 12902289
- PMCID: PMC169109
- DOI: 10.1128/AEM.69.8.4935-4941.2003
Is the in situ accessibility of the 16S rRNA of Escherichia coli for Cy3-labeled oligonucleotide probes predicted by a three-dimensional structure model of the 30S ribosomal subunit?
Abstract
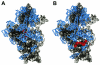
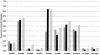
Systematic studies on the hybridization of fluorescently labeled, rRNA-targeted oligonucleotides have shown strong variations in in situ accessibility. Reliable predictions of target site accessibility would contribute to more-rational design of probes for the identification of individual microbial cells in their natural environments. During the past 3 years, numerous studies of the higher-order structure of the ribosome have advanced our understanding of its spatial conformation. These studies range from the identification of rRNA-rRNA interactions based on covariation analyses to physical imaging of the ribosome for the identification of protein-rRNA interactions. Here we reevaluate our Escherichia coli 16S rRNA in situ accessibility data with regard to a tertiary-structure model of the small subunit of the ribosome. We localized target sequences of 176 oligonucleotides on a 3.0-A-resolution three-dimensional (3D) model of the 30S ribosomal subunit. Little correlation was found between probe hybridization efficiency and the proximity of the probe target region to the surface of the 30S ribosomal subunit model. We attribute this to the fact that fluorescence in situ hybridization is performed on fixed cells containing denatured ribosomes, whereas 3D models of the ribosome are based on its native conformation. The effects of different fixation and hybridization protocols on the fluorescence signals conferred by a set of 10 representative probes were tested. The presence or absence of the strongly denaturing detergent sodium dodecyl sulfate had a much more pronounced effect than a change of fixative from paraformaldehyde to ethanol.
Figures



References
-
- Ban, N., P. Nissen, J. Hansen, P. B. Moore, and T. A. Steitz. 2000. The complete atomic structure of the large ribosomal subunit at 2.4 Å resolution. Science 289:905-920. - PubMed
-
- Behrens, S., C. Rühland, J. Inacio, H. Huber, A. Fonseca, I. Spencer-Martins, B. M. Fuchs, and R. Amann. 2003. In situ accessibility of small subunit ribosomal RNA of members of the domains Bacteria, Archaea, and Eucarya to Cy3-labeled oligonucleotide probes. Appl. Environ. Microbiol. 69:1748-1758. - PMC - PubMed
-
- Bogdanov, A. A., N. V. Chichkova, A. M. Kopylov, A. S. Mankin, and E. A. Skripkin. 1988. Surface topography of ribosomal RNA. Methods Enzymol. 164:440-456. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

