The bacterial transposon Tn7 causes premature polyadenylation of mRNA in eukaryotic organisms: TAGKO mutagenesis in filamentous fungi
- PMID: 12907724
- PMCID: PMC169947
- DOI: 10.1093/nar/gkg676
The bacterial transposon Tn7 causes premature polyadenylation of mRNA in eukaryotic organisms: TAGKO mutagenesis in filamentous fungi
Abstract
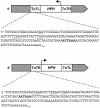
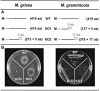
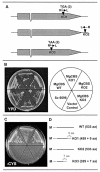
TAGKO is a Tn7-based transposition system for genome wide mutagenesis in filamentous fungi. The effects of transposon insertion on the expression of TAGKO alleles were examined in Magnaporthe grisea and Mycosphaerella graminicola. Northern analysis showed that stable, truncated transcripts were expressed in the TAGKO mutants. Mapping of the 3'-ends of TAGKO cDNAs revealed that they all contain Tn7 end sequences, regardless of the transposon orientation. Polyadenylation signals characteristic of eukaryotic genes, preceded by stop codons in all frames, are located in both ends of the bacterial transposon. Thus, TAGKO transcripts are prematurely polyadenylated, and truncated proteins are predicted to be translated in the fungal mutants. Depending on the extent of protein truncation, TAGKO mutations in HPD4 (encoding p-hydroxyphenylpyruvate dioxygenase) resulted in tyrosine sensitivity in the two fungi. Similarly, a particular M.grisea CBS1 (encoding cystathionine beta-synthase) TAGKO cDNA failed to complement cysteine auxotrophy in a yeast CBS mutant. TAGKO, therefore, represents a useful tool for in vivo study of truncated gene products in filamentous fungi.
Figures

Similar articles
-
Efficient gene identification and targeted gene disruption in the wheat blotch fungus Mycosphaerella graminicola using TAGKO.Curr Genet. 2002 Nov;42(2):123-7. doi: 10.1007/s00294-002-0339-2. Epub 2002 Nov 21. Curr Genet. 2002. PMID: 12478391
-
Gene discovery and gene function assignment in filamentous fungi.Proc Natl Acad Sci U S A. 2001 Apr 24;98(9):5110-5. doi: 10.1073/pnas.091094198. Epub 2001 Apr 10. Proc Natl Acad Sci U S A. 2001. PMID: 11296265 Free PMC article.
-
Molecular characterization of a cystathionine beta-synthase gene, CBS1, in Magnaporthe grisea.Eukaryot Cell. 2002 Apr;1(2):311-4. doi: 10.1128/EC.1.2.311-314.2002. Eukaryot Cell. 2002. PMID: 12455965 Free PMC article.
-
3'-end-forming signals of yeast mRNA.Trends Biochem Sci. 1996 Dec;21(12):477-81. doi: 10.1016/s0968-0004(96)10057-8. Trends Biochem Sci. 1996. PMID: 9009831 Review.
-
Approaches to functional genomics in filamentous fungi.Cell Res. 2006 Jan;16(1):31-44. doi: 10.1038/sj.cr.7310006. Cell Res. 2006. PMID: 16467874 Review.
Cited by
-
Examining the virulence of Candida albicans transcription factor mutants using Galleria mellonella and mouse infection models.Front Microbiol. 2015 May 5;6:367. doi: 10.3389/fmicb.2015.00367. eCollection 2015. Front Microbiol. 2015. PMID: 25999923 Free PMC article.
-
A homologue of the Aspergillus velvet gene regulates both cephalosporin C biosynthesis and hyphal fragmentation in Acremonium chrysogenum.Appl Environ Microbiol. 2007 May;73(10):3412-22. doi: 10.1128/AEM.00129-07. Epub 2007 Mar 30. Appl Environ Microbiol. 2007. PMID: 17400783 Free PMC article.
-
Unexpected Transcripts in Tn7 orf19.2646 C. albicans Mutant Lead to Low Fungal Burden Phenotype In vivo.Front Microbiol. 2017 May 16;8:873. doi: 10.3389/fmicb.2017.00873. eCollection 2017. Front Microbiol. 2017. PMID: 28559890 Free PMC article.
References
-
- McClintok B. (1987) The Discovery and Characterization of Transpososable Elements: the Collected Papers of Barbara McClintok. Garland, New York, NY.
-
- Berg C.M. and Berg,D.E. (1995) Transposable elements as tools for molecular analyses in bacteria. In Sherratt,D.J. (ed.), Mobile Genetics Elements. Oxford University Press, Oxford, UK, pp. 38–68.
-
- Kaiser K., Sentry,J.W. and Finnegan,D.J. (1995) Eukaryotic transposable elements as tools to study gene structure and function. In Sherratt,D.J. (ed.), Mobile Genetics Elements. Oxford University Press, Oxford, UK, pp. 69–100.
-
- Hamer L., DeZwaan,T.M., Montenegro-Chamorro,M.V., Frank,S.A. and Hamer,J.E. (2001) Recent advances in large-scale transposon mutagenesis. Curr. Opin. Chem. Biol., 5, 67–73. - PubMed

