The Arabidopsis NHL3 gene encodes a plasma membrane protein and its overexpression correlates with increased resistance to Pseudomonas syringae pv. tomato DC3000
- PMID: 12913158
- PMCID: PMC181287
- DOI: 10.1104/pp.103.020438
The Arabidopsis NHL3 gene encodes a plasma membrane protein and its overexpression correlates with increased resistance to Pseudomonas syringae pv. tomato DC3000
Abstract
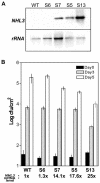
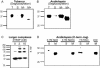
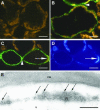
The Arabidopsis genome contains a family of NDR1/HIN1-like (NHL) genes that show homology to the nonrace-specific disease resistance (NDR1) and the tobacco (Nicotiana tabacum) harpin-induced (HIN1) genes. NHL3 is a pathogen-responsive member of this NHL gene family that is potentially involved in defense. In independent transgenic NHL3-overexpressing plant lines, a clear correlation between increased resistance to virulent Pseudomonas syringae pv. tomato DC3000 and enhanced NHL3 transcript levels was seen. These transgenic plants did not show enhanced pathogenesis-related gene expression or reactive oxygen species accumulation. Biochemical and localization experiments were performed to assist elucidation of how NHL3 may confer enhanced disease resistance. Gene constructs expressing amino-terminal c-myc-tagged or carboxyl-terminal hemagglutinin epitope (HA)-tagged NHL3 demonstrated membrane localization in transiently transformed tobacco leaves. Stable Arabidopsis transformants containing the NHL3-HA construct corroborated the findings observed in tobacco. The detected immunoreactive proteins were 10 kD larger than the calculated size and could be partially accounted for by the glycosylation state. However, the expected size was not attained with deglycosylation, suggesting possibly additional posttranslational modification. Detergent treatment, but not chemicals used to strip membrane-associated proteins, could displace the immunoreactive signal from microsomal fractions, showing that NHL3 is tightly membrane associated. Furthermore, immunofluorescence and immunogold labeling, coupled with two-phase partitioning techniques, revealed plasma membrane localization of NHL3-HA. This subcellular localization of NHL3 positions it at an initial contact site to pathogens and may be important in facilitating interception of pathogen-derived signals.
Figures



Similar articles
-
NHL25 and NHL3, two NDR1/HIN1-1ike genes in Arabidopsis thaliana with potential role(s) in plant defense.Mol Plant Microbe Interact. 2002 Jun;15(6):608-16. doi: 10.1094/MPMI.2002.15.6.608. Mol Plant Microbe Interact. 2002. PMID: 12059109
-
The Erwinia amylovora avrRpt2EA gene contributes to virulence on pear and AvrRpt2EA is recognized by Arabidopsis RPS2 when expressed in pseudomonas syringae.Mol Plant Microbe Interact. 2006 Jun;19(6):644-54. doi: 10.1094/MPMI-19-0644. Mol Plant Microbe Interact. 2006. PMID: 16776298
-
Pseudomonas syringae Effector Avirulence Protein E Localizes to the Host Plasma Membrane and Down-Regulates the Expression of the NONRACE-SPECIFIC DISEASE RESISTANCE1/HARPIN-INDUCED1-LIKE13 Gene Required for Antibacterial Immunity in Arabidopsis.Plant Physiol. 2015 Sep;169(1):793-802. doi: 10.1104/pp.15.00547. Epub 2015 Jul 23. Plant Physiol. 2015. PMID: 26206852 Free PMC article.
-
Mimicking the Host Regulation of Salicylic Acid: A Virulence Strategy by the Clubroot Pathogen Plasmodiophora brassicae.Mol Plant Microbe Interact. 2019 Mar;32(3):296-305. doi: 10.1094/MPMI-07-18-0192-R. Epub 2019 Feb 4. Mol Plant Microbe Interact. 2019. PMID: 30199341
-
Characterization of a novel, defense-related Arabidopsis mutant, cir1, isolated by luciferase imaging.Mol Plant Microbe Interact. 2002 Jun;15(6):557-66. doi: 10.1094/MPMI.2002.15.6.557. Mol Plant Microbe Interact. 2002. PMID: 12059104
Cited by
-
An Evolutionarily Conserved DOF-CONSTANS Module Controls Plant Photoperiodic Signaling.Plant Physiol. 2015 Jun;168(2):561-74. doi: 10.1104/pp.15.00321. Epub 2015 Apr 20. Plant Physiol. 2015. PMID: 25897001 Free PMC article.
-
The Soybean Rhg1 locus for resistance to the soybean cyst nematode Heterodera glycines regulates the expression of a large number of stress- and defense-related genes in degenerating feeding cells.Plant Physiol. 2011 Apr;155(4):1960-75. doi: 10.1104/pp.110.167536. Epub 2011 Feb 18. Plant Physiol. 2011. PMID: 21335526 Free PMC article.
-
Transcriptional profiling in response to inhibition of cellulose synthesis by thaxtomin A and isoxaben in Arabidopsis thaliana suspension cells.Plant Cell Rep. 2009 May;28(5):811-30. doi: 10.1007/s00299-009-0670-x. Epub 2009 Feb 7. Plant Cell Rep. 2009. PMID: 19198845
-
Genome-wide analysis of NDR1/HIN1-like genes in pepper (Capsicum annuum L.) and functional characterization of CaNHL4 under biotic and abiotic stresses.Hortic Res. 2020 Jun 1;7(1):93. doi: 10.1038/s41438-020-0318-0. eCollection 2020. Hortic Res. 2020. PMID: 32528705 Free PMC article.
-
Signal molecules mediate the impact of the earthworm Aporrectodea caliginosa on growth, development and defence of the plant Arabidopsis thaliana.PLoS One. 2012;7(12):e49504. doi: 10.1371/journal.pone.0049504. Epub 2012 Dec 3. PLoS One. 2012. PMID: 23226498 Free PMC article.
References
-
- Aoyama T, Chua NH (1997) A glucocorticoid-mediated transcriptional induction system in transgenic plants. Plant J 11: 605–612 - PubMed
-
- Axtell MJ, Staskawicz BJ (2003) Initiation of RPS2-specified disease resistance in Arabidopsis is coupled to the AvrRpt2-directed elimination of RIN4. Cell 112: 369–377 - PubMed
-
- Bonas U, Lahaye T (2001) Plant disease resistance triggered by pathogen-derived molecules: refined models of specific recognition. Curr Opin Microbiol 5: 44–50 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

