HILS1 is a spermatid-specific linker histone H1-like protein implicated in chromatin remodeling during mammalian spermiogenesis
- PMID: 12920187
- PMCID: PMC193598
- DOI: 10.1073/pnas.1837812100
HILS1 is a spermatid-specific linker histone H1-like protein implicated in chromatin remodeling during mammalian spermiogenesis
Abstract
Chromatin remodeling is a major event that occurs during mammalian spermiogenesis, the process of spermatid maturation into spermatozoa. Nuclear condensation during spermiogenesis is accomplished by replacing somatic histones (linker histone H1 and core histones) and the testis-specific linker histone, H1t, with transition proteins and protamines. It has long been thought that H1t is the only testis-specific linker histone, and that all linker histones are replaced by transition proteins, and subsequently by protamines during spermiogenesis. Here, we report the identification and characterization of a spermatid-specific linker histone H1-like protein (termed HILS1) in the mouse and human. Both mouse and human HILS1 genes are located in intron 8 of the alpha-sarcoglycan genes. HILS1 is highly expressed in nuclei of elongating and elongated spermatids (steps 9-15). HILS1 displays several biochemical properties that are similar to those of linker histones, including the abilities to bind reconstituted mononucleosomes, produce a chromatosome stop during micrococcal nuclease digestion, and aggregate chromatin. Because HILS1 is expressed in late spermatids that do not contain core histones, HILS1 may participate in spermatid nuclear condensation through a mechanism distinct from that of linker histones. Because HILS1 also belongs to the large winged helix/forkhead protein superfamily, HILS1 may also regulate gene transcription, DNA repair, and/or other chromosome processes during mammalian spermiogenesis.
Figures


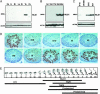


References
-
- Wolffe, A. P. (1997) Int. J. Biochem. Cell Biol. 29, 1463-1466. - PubMed
-
- Hartzog, G. A. & Winston, F. (1997) Curr. Opin. Genet. Dev. 7, 192-198. - PubMed
-
- Wolffe, A. P. (2001) Essays Biochem. 37, 45-57. - PubMed
-
- Lennox, R. W. & Cohen, L. H. (1983) J. Biol. Chem. 258, 262-268. - PubMed
-
- Lennox, R. W. & Cohen, L. H. (1984) Dev. Biol. 103, 80-84. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

