Array analysis of gene expression in connexin-43 null astrocytes
- PMID: 12928503
- PMCID: PMC2651830
- DOI: 10.1152/physiolgenomics.00062.2003
Array analysis of gene expression in connexin-43 null astrocytes
Abstract
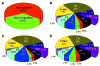
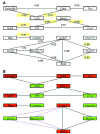
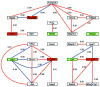
Connexin-43 (Cx43) is the most abundant gap junction protein in brain, where it is found primarily between astrocytes. Although the morphology of astrocytes from Cx43-null (knockout, KO) mice is similar to that of wild-type (WT) astrocytes, KO astrocytes exhibit reduced growth rate in culture. To evaluate the impact of deletion of Cx43 on other genes, including those encoding cell cycle proteins, we used DNA arrays to determine expression patterns in cultured astrocytes from sibling Cx43-null and WT mice. RNA samples extracted from astrocytes cultured from WT and Cx43-null neonatal mice were dye labeled and individually cohybridized with a reference of labeled cDNAs pooled from a variety of tissues on 8 gene arrays containing 8,975 mouse DNA sequences. Normal variability in expression of each gene was evaluated and incorporated into "expression scores" to statistically compare expression levels between WT and KO samples. In Cx43-null astrocytes, 4.1% of the 4,998 adequately quantifiable spots were found to have significantly (P < 0.05) decreased hybridization compared with controls, and 9.4% of the spots showed significantly higher hybridization. The significantly different spots corresponded to RNAs encoding 252 known proteins, many not previously linked to gap junctions, including transcription factors, channels and transporters, cell growth and death signals, enzymes and cell adhesion molecules. These data indicate a surprisingly high degree of impact of deletion of Cx43 on other astrocyte genes, implying that gap junction gene expression alters numerous processes in addition to intercellular communication.
Figures

References
-
- Alberts B, Bray D, Lewis J, Raff M, Roberts K, Watson JD. Molecular Biology of the Cell. 3. New York: Garland; 1994.
-
- Britz-Cunningham SH, Shah MM, Zuppan CW, Fletcher WH. Mutations of the connexin43 gap-junction gene in patients with heart malformations and defects of laterality. N Engl J Med. 1995;332:1323–1329. - PubMed
-
- Butte A. The use and analysis of microarray data. Nat Rev Drug Discov. 2002;1:951–960. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

