Effect of polyamines on the inhibition of peptidyltransferase by antibiotics: revisiting the mechanism of chloramphenicol action
- PMID: 12930958
- PMCID: PMC212793
- DOI: 10.1093/nar/gkg686
Effect of polyamines on the inhibition of peptidyltransferase by antibiotics: revisiting the mechanism of chloramphenicol action
Abstract
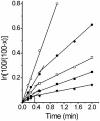
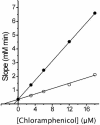
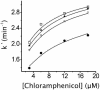
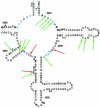
Chloramphenicol is thought to interfere competitively with the binding of the aminoacyl-tRNA 3'-terminus to ribosomal A-site. However, noncompetitive or mixed-noncompetitive inhibition, often observed to be dependent on chloramphenicol concentration and ionic conditions, leaves some doubt about the precise mode of action. Here, we examine further the inhibition effect of chloramphenicol, using a model system derived from Escherichia coli in which a peptide bond is formed between puromycin and AcPhe-tRNA bound at the P-site of poly(U)-programmed ribosomes, under ionic conditions (6 mM Mg2+, 100 mM NH4+, 100 microM spermine) more closely resembling the physiological status. Kinetics reveal that chloramphenicol (I) reacts rapidly with AcPhe-tRNA.poly(U).70S ribosomal complex (C) to form the encounter complex CI which is then isomerized slowly to a more tight complex, C*I. A similar inhibition pattern is observed, if complex C modified by a photoreactive analogue of spermine, reacts in buffer free of spermine. Spermine, either reversibly interacting with or covalently attached to ribosomes, enhances the peptidyltransferase activity and increases the chloramphenicol potency, without affecting the isomerization step. As indicated by photoaffinity labeling, the peptidyltransferase center at which chloramphenicol binds, is one of the preferred cross-linking sites for polyamines. This fact may explain the effect of spermine on chloramphenicol binding to ribosomes.
Figures



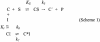
Similar articles
-
Localization of spermine binding sites in 23S rRNA by photoaffinity labeling: parsing the spermine contribution to ribosomal 50S subunit functions.Nucleic Acids Res. 2005 May 16;33(9):2792-805. doi: 10.1093/nar/gki557. Print 2005. Nucleic Acids Res. 2005. PMID: 15897324 Free PMC article.
-
Effects of two photoreactive spermine analogues on peptide bond formation and their application for labeling proteins in Escherichia coli functional ribosomal complexes.Biochemistry. 2001 Jun 26;40(25):7641-50. doi: 10.1021/bi010010s. Biochemistry. 2001. PMID: 11412118
-
Photoaffinity polyamines: interactions with AcPhe-tRNA free in solution or bound at the P-site of Escherichia coli ribosomes.Nucleic Acids Res. 2000 Oct 1;28(19):3733-42. doi: 10.1093/nar/28.19.3733. Nucleic Acids Res. 2000. PMID: 11000265 Free PMC article.
-
Movement of the 3'-end of tRNA through the peptidyl transferase centre and its inhibition by antibiotics.FEBS Lett. 1997 Apr 14;406(3):223-33. doi: 10.1016/s0014-5793(97)00261-5. FEBS Lett. 1997. PMID: 9136892 Review.
-
The molecular basis of the inhibitory activities of type A and type B synergimycins and related antibiotics on ribosomes.J Antimicrob Chemother. 1989 Oct;24(4):485-507. doi: 10.1093/jac/24.4.485. J Antimicrob Chemother. 1989. PMID: 2515187 Review.
Cited by
-
Chloramphenicol Derivatives as Antibacterial and Anticancer Agents: Historic Problems and Current Solutions.Antibiotics (Basel). 2016 Jun 3;5(2):20. doi: 10.3390/antibiotics5020020. Antibiotics (Basel). 2016. PMID: 27271676 Free PMC article. Review.
-
Revisiting the structures of several antibiotics bound to the bacterial ribosome.Proc Natl Acad Sci U S A. 2010 Oct 5;107(40):17158-63. doi: 10.1073/pnas.1008685107. Epub 2010 Sep 27. Proc Natl Acad Sci U S A. 2010. PMID: 20876130 Free PMC article.
-
Conjugation with polyamines enhances the antibacterial and anticancer activity of chloramphenicol.Nucleic Acids Res. 2014 Jul;42(13):8621-34. doi: 10.1093/nar/gku539. Epub 2014 Jun 17. Nucleic Acids Res. 2014. PMID: 24939899 Free PMC article.
-
Synthesis and evaluation of chloramphenicol homodimers: molecular target, antimicrobial activity, and toxicity against human cells.PLoS One. 2015 Aug 12;10(8):e0134526. doi: 10.1371/journal.pone.0134526. eCollection 2015. PLoS One. 2015. PMID: 26267355 Free PMC article.
-
Visualizing the modification landscape of the human 60S ribosomal subunit at close to atomic resolution.Nucleic Acids Res. 2025 Jan 7;53(1):gkae1191. doi: 10.1093/nar/gkae1191. Nucleic Acids Res. 2025. PMID: 39658079 Free PMC article.
References
-
- Wareham D.W. and Wilson,P. (2002) Chloramphenicol in the 21st century. Hosp. Med., 63, 157–161. - PubMed
-
- Pestka S. (1970) Formation of transfer ribonucleic acid-ribosome complexes. VII. Survey of the effect of antibiotics on N-acetylphenylalanyl puromycin formation: possible mechanism of chloramphenicol action. Arch. Biochem. Biophys., 136, 80–88. - PubMed
-
- Fernandez-Muñoz R. and Vazquez,D. (1973) Kinetic studies of peptide bond formation. Effect of chloramphenicol. Mol. Biol. Rep., 1, 75–79. - PubMed
-
- Bhuta P., Chung,H.L., Hwang,J.-S. and Žemlička,J. (1980) Analogues of chloramphenicol: circular dichroism spectra, inhibition of ribosomal peptidyltransferase, and possible mechanism of action. J. Med. Chem., 23, 1299–1305. - PubMed
-
- Drainas D., Kalpaxis,D.L. and Coutsogeorgopoulos,C. (1987) Inhibition of ribosomal peptidyltransferase by chloramphenicol: kinetic studies. Eur. J. Biochem., 164, 53–68. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

