Phosphohydrolase and transphosphatidylation reactions of two Streptomyces phospholipase D enzymes: covalent versus noncovalent catalysis
- PMID: 12931007
- PMCID: PMC2324005
- DOI: 10.1110/ps.03192503
Phosphohydrolase and transphosphatidylation reactions of two Streptomyces phospholipase D enzymes: covalent versus noncovalent catalysis
Abstract
A kinetic comparison of the hydrolase and transferase activities of two bacterial phospholipase D (PLD) enzymes with little sequence homology provides insights into mechanistic differences and also the more general role of Ca(2+) in modulating PLD reactions. Although the two PLDs exhibit similar substrate specificity (phosphatidylcholine preferred), sensitivity to substrate aggregation or Ca(2+), and pH optima are quite distinct. Streptomyces sp. PMF PLD, a member of the PLD superfamily, generates both hydrolase and transferase products in parallel, consistent with a mechanism that proceeds through a covalent phosphatidylhistidyl intermediate where the rate-limiting step is formation of the covalent intermediate. For Streptomyces chromofuscus PLD, the two reactions exhibit different pH profiles, a result consistent with a mechanism likely to involve direct attack of water or an alcohol on the phosphorus. Ca(2+), not required for monomer or micelle hydrolysis, can activate both PLDs for hydrolysis of PC unilamellar vesicles. In the case of Streptomyces sp. PMF PLD, Ca(2+) relieves product inhibition by interactions with the phosphatidic acid (PA). A similar rate enhancement could occur with other HxKx(4)D-motif PLDs as well. For S. chromofuscus PLD, Ca(2+) is absolutely critical for binding of the enzyme to PC vesicles and for PA activation. That the Ca(2+)-PA activation involves a discreet site on the protein is suggested by the observation that the identity of the C-terminal residue in S. chromofuscus PLD can modulate the extent of product activation.
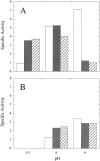
Figures
Similar articles
-
The role of interfacial binding in the activation of Streptomyces chromofuscus phospholipase D by phosphatidic acid.J Biol Chem. 1999 Dec 10;274(50):35367-74. doi: 10.1074/jbc.274.50.35367. J Biol Chem. 1999. PMID: 10585404
-
Binding of proteolytically processed phospholipase D from Streptomyces chromofuscus to phosphatidylcholine membranes facilitates vesicle aggregation and fusion.Biochemistry. 2001 Nov 20;40(46):13954-63. doi: 10.1021/bi011338o. Biochemistry. 2001. PMID: 11705386
-
Expression and characterization of a heterodimer of Streptomyces chromofuscus phospholipase D.Biochim Biophys Acta. 2004 Dec 1;1703(1):43-51. doi: 10.1016/j.bbapap.2004.09.014. Biochim Biophys Acta. 2004. PMID: 15588701
-
Phospholipase D mechanism using Streptomyces PLD.Biochim Biophys Acta. 2009 Sep;1791(9):962-9. doi: 10.1016/j.bbalip.2009.01.020. Epub 2009 Feb 4. Biochim Biophys Acta. 2009. PMID: 19416643 Review.
-
Phospholipase D and its application in biocatalysis.Biotechnol Lett. 2005 Apr;27(8):535-44. doi: 10.1007/s10529-005-3251-2. Biotechnol Lett. 2005. PMID: 15973486 Review.
Cited by
-
Phospholipase D activity is regulated by product segregation and the structure formation of phosphatidic acid within model membranes.Biophys J. 2007 Oct 1;93(7):2373-83. doi: 10.1529/biophysj.107.108787. Epub 2007 Jun 8. Biophys J. 2007. PMID: 17557794 Free PMC article.
-
Molecular cloning, heterologous expression, and enzymatic characterization of lysoplasmalogen-specific phospholipase D from Thermocrispum sp.FEBS Open Bio. 2016 Oct 17;6(11):1113-1130. doi: 10.1002/2211-5463.12131. eCollection 2016 Nov. FEBS Open Bio. 2016. PMID: 27833852 Free PMC article.
-
A Novel High-Throughput Assay Reveals That the Temperature Induced Increases in Transphosphatidylation of Phospholipase D Are Dependent on the Alcohol Acceptor Concentration.Biomolecules. 2022 Apr 25;12(5):632. doi: 10.3390/biom12050632. Biomolecules. 2022. PMID: 35625563 Free PMC article.
-
Ultralow background membrane editors for spatiotemporal control of lipid metabolism and signaling.bioRxiv [Preprint]. 2023 Aug 31:2023.08.31.555787. doi: 10.1101/2023.08.31.555787. bioRxiv. 2023. Update in: ACS Cent Sci. 2024 Jan 30;10(3):543-554. doi: 10.1021/acscentsci.3c01105. PMID: 37693485 Free PMC article. Updated. Preprint.
-
Ultralow Background Membrane Editors for Spatiotemporal Control of Phosphatidic Acid Metabolism and Signaling.ACS Cent Sci. 2024 Jan 30;10(3):543-554. doi: 10.1021/acscentsci.3c01105. eCollection 2024 Mar 27. ACS Cent Sci. 2024. PMID: 38559292 Free PMC article.
References
-
- Bian, J. and Roberts, M.F. 1992. Thermodynamic comparison of lyso- and diacylphosphatidylcholines. J. Colloid Interface Sci. 153 420–428.
-
- Dorovska-Taran, V., Wick, R., and Walde, P. 1996. A 1H nuclear magnetic resonance method for investigating the phosholipase D-catalyzed hydrolysis of phosphatidylcholine in liposomes. Anal. Biochem. 240 37–47. - PubMed
-
- Eibl, H. and Kovatchev, S. 1981. Preparation of phospholipids and their analogues by phospholipase D. Methods Enzymol. 72 632–639. - PubMed
-
- El Kirat, K., Besson, F., Prigent, A.F., Chauvet, J.P., and Roux, B. 2002. Role of calcium and membrane organization on phospholipase D localization and activity. Competition between a soluble and insoluble substrate. J. Biol. Chem. 277 21231–21236. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

