Asymmetric mutation rates at enzyme-inhibitor interfaces: implications for the protein-protein docking problem
- PMID: 12931008
- PMCID: PMC2324006
- DOI: 10.1110/ps.0306303
Asymmetric mutation rates at enzyme-inhibitor interfaces: implications for the protein-protein docking problem
Abstract
We have carried out a thorough and systematic sequence-structure study on how the pattern of conservation at the interface differs from the noninteracting surface in seven proteases and their inhibitors. As expected, the interface of a protease could be easily distinguished from the noninteracting surface by a concentrated area of conservation. In contrast, there was less distinction to be made between the interface and the noninteracting surface of inhibitors, and in five of the seven cases, a higher proportion of the interface area was variable compared to the rest of the surface. This is likely to cause a problem for binding-site prediction methods that assume the largest cluster of highly conserved residues on the surface of a protein corresponds to the interface. We conclude that such methods would succeed when applied to our protease test cases, but complications could arise with the inhibitors. These results also impact on methods to solve the protein-protein docking problem that use conservation at the interface to provide the location of the two protein binding sites prior to application of the docking algorithm.
Figures
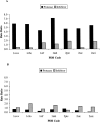
Similar articles
-
Looking at enzymes from the inside out: the proximity of catalytic residues to the molecular centroid can be used for detection of active sites and enzyme-ligand interfaces.J Mol Biol. 2005 Aug 12;351(2):309-26. doi: 10.1016/j.jmb.2005.06.047. J Mol Biol. 2005. PMID: 16019028
-
Docking of protein molecular surfaces with evolutionary trace analysis.Proteins. 2007 Dec 1;69(4):832-8. doi: 10.1002/prot.21737. Proteins. 2007. PMID: 17803239
-
Are protein-protein interfaces more conserved in sequence than the rest of the protein surface?Protein Sci. 2004 Jan;13(1):190-202. doi: 10.1110/ps.03323604. Protein Sci. 2004. PMID: 14691234 Free PMC article.
-
Comparing protein-ligand docking programs is difficult.Proteins. 2005 Aug 15;60(3):325-32. doi: 10.1002/prot.20497. Proteins. 2005. PMID: 15937897 Review.
-
Interpretive proteomics--finding biological meaning in genome and proteome databases.Adv Enzyme Regul. 2003;43:271-359. doi: 10.1016/s0065-2571(02)00024-9. Adv Enzyme Regul. 2003. PMID: 12791396 Review. No abstract available.
Cited by
-
Protein-protein binding supersites.PLoS Comput Biol. 2019 Jan 7;15(1):e1006704. doi: 10.1371/journal.pcbi.1006704. eCollection 2019 Jan. PLoS Comput Biol. 2019. PMID: 30615604 Free PMC article.
-
Algorithmic approaches to protein-protein interaction site prediction.Algorithms Mol Biol. 2015 Feb 15;10:7. doi: 10.1186/s13015-015-0033-9. eCollection 2015. Algorithms Mol Biol. 2015. PMID: 25713596 Free PMC article.
-
Joint evolutionary trees: a large-scale method to predict protein interfaces based on sequence sampling.PLoS Comput Biol. 2009 Jan;5(1):e1000267. doi: 10.1371/journal.pcbi.1000267. Epub 2009 Jan 23. PLoS Comput Biol. 2009. PMID: 19165315 Free PMC article.
-
Protein binding site prediction using an empirical scoring function.Nucleic Acids Res. 2006 Aug 7;34(13):3698-707. doi: 10.1093/nar/gkl454. Print 2006. Nucleic Acids Res. 2006. PMID: 16893954 Free PMC article.
References
-
- Beuning, L.L., Spriggs, T.W., and Christeller, J.T. 1994. Evolution of the proteinase inhibitor I family and apparent lack of hypervariability in the proteinase contact group. J. Mol. Evol. 39 644–654. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

