Characterization of Pseudomonas aeruginosa exoenzyme S as a bifunctional enzyme in J774A.1 macrophages
- PMID: 12933877
- PMCID: PMC187317
- DOI: 10.1128/IAI.71.9.5296-5305.2003
Characterization of Pseudomonas aeruginosa exoenzyme S as a bifunctional enzyme in J774A.1 macrophages
Abstract
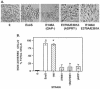
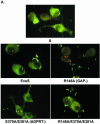
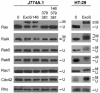
Pseudomonas aeruginosa exoenzyme S (ExoS) is a type III secretion (TTS) effector, which includes both a GTPase-activating protein (GAP) activity toward the Rho family of low-molecular-weight G (LMWG) proteins and an ADP-ribosyltransferase (ADPRT) activity that targets LMWG proteins in the Ras, Rab, and Rho families. The coordinate function of both activities of ExoS in J774A.1 macrophages was assessed by using P. aeruginosa strains expressing and translocating wild-type ExoS or ExoS defective in GAP and/or ADPRT activity. Distinct and coordinated functions were identified for both domains. The GAP activity was required for the antiphagocytic effect of ExoS and was linked to interference of lamellopodium and membrane ruffle formation. Alternatively, the ADPRT activity of ExoS altered cellular adherence and morphology and was linked to effects on filopodium formation. The cellular mechanism of ExoS GAP activity included an inactivation of Rac1 function, as determined in p21-activated kinase 1-glutathione S-transferase (GST) pull-down assays. The ADPRT activity of ExoS targeted Ras and RalA but not Rab or Rho proteins, and Ral binding protein 1-GST pull-down assays identified an effect of ExoS ADPRT activity on RalA activation. The results from these studies confirm the bifunctional nature of ExoS activity within macrophages when translocated by TTS.
Figures



References
-
- Aktories, K., C. Mohr, and G. Koch. 1992. Clostridium botulinum C3 ADP-ribosyltransferase. Curr. Top. Microbiol. Immunol. 175:115-131. - PubMed
-
- Allen, W. E., G. E. Jones, J. W. Pollard, and A. J. Ridley. 1997. Rho, Rac and Cdc42 regulate actin organization and cell adhesion in macrophages. J. Cell Sci. 110:707-720. - PubMed
-
- Caron, E., and A. Hall. 1998. Identification of two distinct mechanisms of phagocytosis controlled by different Rho GTPases. Science 282:1717-1721. - PubMed
-
- Coburn, J., R. T. Wyatt, B. H. Iglewski, and D. M. Gill. 1989. Several GTP-binding proteins, including p21c-H-ras, are preferred substrates of Pseudomonas aeruginosa exoenzyme S. J. Biol. Chem. 264:9004-9008. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

