Full-length archaeal Rad51 structure and mutants: mechanisms for RAD51 assembly and control by BRCA2
- PMID: 12941707
- PMCID: PMC202371
- DOI: 10.1093/emboj/cdg429
Full-length archaeal Rad51 structure and mutants: mechanisms for RAD51 assembly and control by BRCA2
Abstract
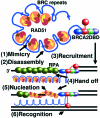
To clarify RAD51 interactions controlling homologous recombination, we report here the crystal structure of the full-length RAD51 homolog from Pyrococcus furiosus. The structure reveals how RAD51 proteins assemble into inactive heptameric rings and active DNA-bound filaments matching three-dimensional electron microscopy reconstructions. A polymerization motif (RAD51-PM) tethers individual subunits together to form assemblies. Subunit interactions support an allosteric 'switch' promoting ATPase activity and DNA binding roles for the N-terminal domain helix-hairpin-helix (HhH) motif. Structural and mutational results characterize RAD51 interactions with the breast cancer susceptibility protein BRCA2 in higher eukaryotes. A designed P.furiosus RAD51 mutant binds BRC repeats and forms BRCA2-dependent nuclear foci in human cells in response to gamma-irradiation-induced DNA damage, similar to human RAD51. These results show that BRCA2 repeats mimic the RAD51-PM and imply analogous RAD51 interactions with RAD52 and RAD54. Both BRCA2 and RAD54 may act as antagonists and chaperones for RAD51 filament assembly by coupling RAD51 interface exchanges with DNA binding. Together, these structural and mutational results support an interface exchange hypothesis for coordinated protein interactions in homologous recombination.
Figures





Similar articles
-
The rad50 signature motif: essential to ATP binding and biological function.J Mol Biol. 2004 Jan 23;335(4):937-51. doi: 10.1016/j.jmb.2003.11.026. J Mol Biol. 2004. PMID: 14698290
-
BRCA2 C-terminal clamp restructures RAD51 dimers to bind B-DNA for replication fork stability.Mol Cell. 2025 Jun 5;85(11):2080-2096.e6. doi: 10.1016/j.molcel.2025.05.010. Epub 2025 May 28. Mol Cell. 2025. PMID: 40441151
-
A large C-terminal Rad52 segment acts as a chaperone to Form and Stabilize Rad51 Filaments.Nat Commun. 2025 Jul 1;16(1):5589. doi: 10.1038/s41467-025-60664-x. Nat Commun. 2025. PMID: 40595518 Free PMC article.
-
Overexpression of RAD51 suppresses recombination defects: a possible mechanism to reverse genomic instability.Nucleic Acids Res. 2010 Mar;38(4):1061-70. doi: 10.1093/nar/gkp1063. Epub 2009 Nov 26. Nucleic Acids Res. 2010. PMID: 19942681 Free PMC article. Review.
-
Structural insights into BRCA2 function.Curr Opin Struct Biol. 2003 Apr;13(2):206-11. doi: 10.1016/s0959-440x(03)00033-2. Curr Opin Struct Biol. 2003. PMID: 12727514 Review.
Cited by
-
A new protein complex promoting the assembly of Rad51 filaments.Nat Commun. 2013;4:1676. doi: 10.1038/ncomms2678. Nat Commun. 2013. PMID: 23575680 Free PMC article.
-
Identifying DNA-binding proteins using structural motifs and the electrostatic potential.Nucleic Acids Res. 2004 Sep 8;32(16):4732-41. doi: 10.1093/nar/gkh803. Print 2004. Nucleic Acids Res. 2004. PMID: 15356290 Free PMC article.
-
Cells expressing murine RAD52 splice variants favor sister chromatid repair.Mol Cell Biol. 2006 May;26(10):3752-63. doi: 10.1128/MCB.26.10.3752-3763.2006. Mol Cell Biol. 2006. PMID: 16648471 Free PMC article.
-
An archaeal Rad54 protein remodels DNA and stimulates DNA strand exchange by RadA.Nucleic Acids Res. 2009 May;37(8):2757-70. doi: 10.1093/nar/gkp068. Epub 2009 Mar 12. Nucleic Acids Res. 2009. PMID: 19282450 Free PMC article.
-
BRCA-associated ovarian cancer: from molecular genetics to risk management.Biomed Res Int. 2014;2014:787143. doi: 10.1155/2014/787143. Epub 2014 Jul 22. Biomed Res Int. 2014. PMID: 25136623 Free PMC article. Review.
References
-
- Aihara H., Ito,Y., Kurumizaka,H., Terada,T., Yokoyama,S. and Shibata,T. (1997) An interaction between a specified surface of the C-terminal domain of RecA protein and double-stranded DNA for homologous pairing. J. Mol. Biol., 274, 213–221. - PubMed
-
- Aihara H., Ito,Y., Kurumizaka,H., Yokoyama,S. and Shibata,T. (1999) The N-terminal domain of the human Rad51 protein binds DNA: structure and a DNA binding surface as revealed by NMR. J. Mol. Biol., 290, 495–504. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

