Similarity of force-induced unfolding of apomyoglobin to its chemical-induced unfolding: an atomistic molecular dynamics simulation approach
- PMID: 12944267
- PMCID: PMC1303326
- DOI: 10.1016/S0006-3495(03)74582-2
Similarity of force-induced unfolding of apomyoglobin to its chemical-induced unfolding: an atomistic molecular dynamics simulation approach
Abstract
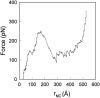
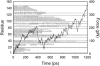
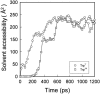
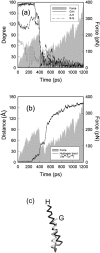
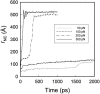
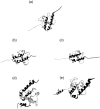
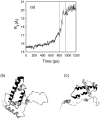
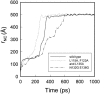
We have compared force-induced unfolding with traditional unfolding methods using apomyoglobin as a model protein. Using molecular dynamics simulation, we have investigated the structural stability as a function of the degree of mechanical perturbation. Both anisotropic perturbation by stretching two terminal atoms and isotropic perturbation by increasing the radius of gyration of the protein show the same key event of force-induced unfolding. Our primary results show that the native structure of apomyoglobin becomes destabilized against the mechanical perturbation as soon as the interhelical packing between the G and H helices is broken, suggesting that our simulation results share a common feature with the experimental observation that the interhelical contact is more important for the folding of apomyoglobin than the stability of individual helices. This finding is further confirmed by simulating both helix destabilizing and interhelical packing destabilizing mutants.
Figures


Similar articles
-
Cooperativity of folding of the apomyoglobin pH 4 intermediate studied by glycine and proline mutations.Nat Struct Biol. 1997 Nov;4(11):925-30. doi: 10.1038/nsb1197-925. Nat Struct Biol. 1997. PMID: 9360609
-
Molten globular characteristics of the native state of apomyoglobin.Nat Struct Biol. 1994 Jul;1(7):447-52. doi: 10.1038/nsb0794-447. Nat Struct Biol. 1994. PMID: 7664063
-
Packing interactions in the apomyglobin folding intermediate.Nat Struct Biol. 1996 May;3(5):439-45. doi: 10.1038/nsb0596-439. Nat Struct Biol. 1996. PMID: 8612074
-
The folding process of apomyoglobin.Protein Pept Lett. 2005 Apr;12(3):229-34. doi: 10.2174/0929866053587174. Protein Pept Lett. 2005. PMID: 15777270 Review.
-
Misfolding and amyloid aggregation of apomyoglobin.Int J Mol Sci. 2013 Jul 9;14(7):14287-300. doi: 10.3390/ijms140714287. Int J Mol Sci. 2013. PMID: 23839096 Free PMC article. Review.
Cited by
-
Application of conventional molecular dynamics simulation in evaluating the stability of apomyoglobin in urea solution.Sci Rep. 2017 Mar 16;7:44651. doi: 10.1038/srep44651. Sci Rep. 2017. PMID: 28300210 Free PMC article.
-
A Computational Approach to Modeling an Antagonistic Angiogenic VEGFR1-IL2 Fusion Protein for Cancer Therapy.Bioinform Biol Insights. 2021 Sep 20;15:11779322211043297. doi: 10.1177/11779322211043297. eCollection 2021. Bioinform Biol Insights. 2021. PMID: 34566410 Free PMC article.
References
-
- Barrick, D., and R. L. Baldwin. 1993. Three-state analysis of sperm whale apomyoglobin folding. Biochemistry. 32:3790–3796. - PubMed
-
- Brooks, B. R., R. E. Bruccoleri, B. D. Olafson, D. J. States, S. Swaminathan, and M. Karplus. 1983. CHARMM: a program for macromolecular energy, minimization and dynamics calculations. J. Comput. Chem. 4:187–217.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

