Multiplexed DNA quantification by spectroscopic shift of two microsphere cavities
- PMID: 12944310
- PMCID: PMC1303369
- DOI: 10.1016/S0006-3495(03)74625-6
Multiplexed DNA quantification by spectroscopic shift of two microsphere cavities
Abstract
We have developed a novel, spectroscopic technique for high-sensitivity, label-free DNA quantification. We demonstrate that an optical resonance (whispering gallery mode) excited in a micron-sized silica sphere can be used to detect and measure nucleic acids. The surface of the silica sphere is chemically modified with oligonucleotides. We show that hybridization to the target DNA leads to a red shift of the optical resonance wavelength. The sensitivity of this resonant technique is measured as 6 pg/mm(2) mass loading, higher as compared to most optical single-pass devices such as surface plasmon resonance biosensors. Furthermore, we show that each microsphere can be identified by its unique resonance wavelength. Specific, multiplexed DNA detection is demonstrated by using two microspheres. The multiplexed signal from two microspheres allows us to discriminate a single nucleotide mismatch in an 11-mer oligonucleotide with a high signal-to-noise ratio of 54. This all-photonic whispering gallery mode biosensor can be integrated on a semiconductor chip that makes it an easy to manufacture, analytic component for a portable, robust lab-on-a-chip device.
Figures



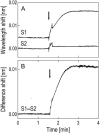
References
-
- Armani, D. K., T. J. Kippenberg, S. M. Spillane, and K. J. Vahala. 2003. Ultra-high-Q toroid microcavity on a chip. Nature. 421:925–928. - PubMed
-
- Arnold, S., M. Khoshsima, I. Teraoka, S. Holler, and F. Vollmer. 2003. Shift of whispering gallery modes in microspheres by protein adsorption. Opt. Lett. 28:272–274. - PubMed
-
- Baird, C. L., and D. G. Myszka. 2001. Current and emerging commercial optical biosensors. J. Mol. Recognit. 14:261–268. - PubMed
-
- Bates, P. J., J. F. Reddoch, P. Hansakul, A. Arrow, R. Dale, and D. M. Miller. 2002. Biosensor detection of triplex formation by modified oligonucleotides. Anal. Biochem. 307:235–243. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

