VirE2, a type IV secretion substrate, interacts with the VirD4 transfer protein at cell poles of Agrobacterium tumefaciens
- PMID: 12950931
- PMCID: PMC3882298
- DOI: 10.1046/j.1365-2958.2003.03669.x
VirE2, a type IV secretion substrate, interacts with the VirD4 transfer protein at cell poles of Agrobacterium tumefaciens
Abstract
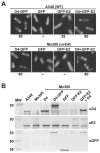
Agrobacterium tumefaciens transfers oncogenic DNA and effector proteins to plant cells during the course of infection. Substrate translocation across the bacterial cell envelope is mediated by a type IV secretion (TFS) system composed of the VirB proteins, as well as VirD4, a member of a large family of inner membrane proteins implicated in the coupling of DNA transfer intermediates to the secretion machine. In this study, we demonstrate with novel cytological screens - a two-hybrid (C2H) assay and bimolecular fluorescence complementation (BiFC) - and by immunoprecipitation of chemically cross-linked protein complexes that the VirE2 effector protein interacts directly with the VirD4 coupling protein at cell poles of A. tumefaciens. Analyses of truncation derivatives showed that VirE2 interacts via its C terminus with VirD4, and, further, an NH2-terminal membrane-spanning domain of VirD4 is dispensable for complex formation. VirE2 interacts with VirD4 independently of the virB-encoded transfer machine and T pilus, the putative periplasmic chaperones AcvB and VirJ, and the T-DNA transfer intermediate. Finally, VirE2 is recruited to polar-localized VirD4 as a complex with its stabilizing secretion chaperone VirE1, yet the effector-coupling protein interaction is not dependent on chaperone binding. Together, our findings establish for the first time that a protein substrate of a type IV secretion system is recruited to a member of the coupling protein superfamily.
Figures

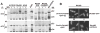


References
-
- Backert S, Ziska E, Brinkmann V, Zimny-Arndt U, Fauconnier A, Jungblut PR, et al. Translocation of the Helicobacter pylori CagA protein in gastric epithelial cells by a type IV secretion apparatus. Cell Microbiol. 2000;2:155–164. - PubMed
-
- Baron C, O’Callaghan D, Lanka E. Bacterial secrets of secretion: Euro Conference on the biology of type IV secretion processes. Mol Microbiol. 2002;43:1359–1365. - PubMed
-
- Burgess RR, Arthur TM, Pietz BC. Mapping protein–protein interaction domains using ordered fragment ladder far-western analysis of hexahistidine-tagged fusion proteins. Methods Enzymol. 2000;328:141–157. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

