Pleiotropy, homeostasis, and functional networks based on assays of cardiovascular traits in genetically randomized populations
- PMID: 12952877
- PMCID: PMC403692
- DOI: 10.1101/gr.1186603
Pleiotropy, homeostasis, and functional networks based on assays of cardiovascular traits in genetically randomized populations
Abstract
A major problem in studying biological traits is understanding how genes work together to provide organismal structures and functions. Conventional reductionist paradigms attribute functions to particular proteins, motifs, and amino acids. An equally important but harder problem involves the synthesis of data at fundamental levels of biological systems to understand functionality at higher levels. We used subtle, naturally occurring, multigenic variation of cardiovascular (CV) properties in a panel of genetically randomized strains that are derived from the A/J and C57BL/6J strains of mice to perturb CV functions in nonpathologic ways. In this proof-of-concept study, computational analysis correctly identified the known relations among CV properties and revealed functionality at higher levels of the CV system. The network was then used to account for pleiotropies and homeostatic responses in single gene mutant mice and in mice treated with a pharmacologic agent (anesthesia). The CV network accounted for functional dependencies in complementary ways to the insights obtained from genetic networks and biochemical pathways. These networks are therefore an important approach for defining and characterizing functional relations in complex biological systems in health and disease.
Figures



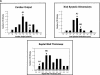







References
-
- Alonso-Blanco, C., Koornneef, M., and Stam, P. 1998. The use of recombinant inbred lines (RILs) for genetic mapping. Methods Mol. Biol. 82: 137–146. - PubMed
-
- Arkin, A., Shen, P., and Ross, J. 1997. A test case for correlation metric construction of a reaction pathway from measurements. Science 277: 1275–1279.
-
- ____. 1981. Recombinant inbred strains and bilineal congenic strains. In The mouse in biomedical research, Vol. 1. History, genetics and wild mice (eds. H.L. Foster et al.), pp. 223–239. Academic Press, New York.
-
- Brem, R.B., Yvert, G., Clinton, R., and Kruglyak, L. 2002. Genetic dissection of transcriptional regulation in budding yeast. Science 296: 752–755. - PubMed
WEB SITE REFERENCES
-
- http://www.jax.org/staff/churchill/labsite/dataset/index.html; Raw data in tab delineated format along with a script that can be executed in a Matlab (Mathworks Inc., Natick, MA) session to replicate the analyses described in this article.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
