Identification of the receptor binding domain of the mouse mammary tumor virus envelope protein
- PMID: 12970432
- PMCID: PMC228533
- DOI: 10.1128/jvi.77.19.10468-10478.2003
Identification of the receptor binding domain of the mouse mammary tumor virus envelope protein
Abstract
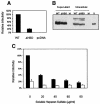
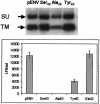
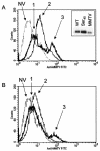
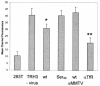
Mouse mammary tumor virus (MMTV) is a betaretrovirus that infects rodent cells and uses mouse transferrin receptor 1 for cell entry. To characterize the interaction of MMTV with its receptor, we aligned the MMTV envelope surface (SU) protein with that of Friend murine leukemia virus (F-MLV) and identified a putative receptor-binding domain (RBD) that included a receptor binding sequence (RBS) of five amino acids and a heparin-binding domain (HBD). Mutation of the HBD reduced virus infectivity, and soluble heparan sulfate blocked infection of cells by wild-type pseudovirus. Interestingly, some but not all MMTV-like elements found in primary and cultured human breast cancer cell lines, termed h-MTVs, had sequence alterations in the putative RBS. Single substitution of one of the amino acids found in an h-MTV RBS variant in the RBD of MMTV, Phe(40) to Ser, did not alter species tropism but abolished both virus binding to cells and infectivity. Neutralizing anti-SU monoclonal antibodies also recognized a glutathione S-transferase fusion protein that contained the five-amino-acid RBS region from MMTV. The critical Phe(40) residue is located on a surface of the MMTV RBD model that is distant from and may be structurally more rigid than the region of F-MLV RBD that contains its critical binding site residues. This suggests that, in contrast to other murine retroviruses, binding to its receptor may result in few or no changes in MMTV envelope protein conformation.
Figures



Similar articles
-
Single amino acid substitution (G42E) in the receptor binding domain of mouse mammary tumour virus envelope protein facilitates infection of non-murine cells in a transferrin receptor 1-independent manner.Retrovirology. 2015 May 16;12:43. doi: 10.1186/s12977-015-0168-2. Retrovirology. 2015. PMID: 25980759 Free PMC article.
-
Envelope proteins containing single amino acid substitutions support a structural model of the receptor-binding domain of bovine leukemia virus surface protein.J Virol. 2002 Nov;76(21):10861-72. doi: 10.1128/jvi.76.21.10861-10872.2002. J Virol. 2002. PMID: 12368329 Free PMC article.
-
Mouse transferrin receptor 1 is the cell entry receptor for mouse mammary tumor virus.Proc Natl Acad Sci U S A. 2002 Sep 17;99(19):12386-90. doi: 10.1073/pnas.192360099. Epub 2002 Sep 6. Proc Natl Acad Sci U S A. 2002. PMID: 12218182 Free PMC article.
-
Mouse Mammary Tumour Virus (MMTV) in Human Breast Cancer-The Value of Bradford Hill Criteria.Viruses. 2022 Mar 30;14(4):721. doi: 10.3390/v14040721. Viruses. 2022. PMID: 35458452 Free PMC article. Review.
-
Involvement of a mouse mammary tumor virus (MMTV) homologue in human breast cancer: Evidence for, against and possible causes of controversies.Microb Pathog. 2019 May;130:283-294. doi: 10.1016/j.micpath.2019.03.021. Epub 2019 Mar 21. Microb Pathog. 2019. PMID: 30905715 Review.
Cited by
-
Substitutions of conserved amino acids in the receptor-binding domain of the spike glycoprotein affect utilization of murine CEACAM1a by the murine coronavirus MHV-A59.Virology. 2005 Mar 30;334(1):98-110. doi: 10.1016/j.virol.2005.01.016. Virology. 2005. PMID: 15749126 Free PMC article.
-
Domain Organization of Lentiviral and Betaretroviral Surface Envelope Glycoproteins Modeled with AlphaFold.J Virol. 2022 Jan 26;96(2):e0134821. doi: 10.1128/JVI.01348-21. Epub 2021 Oct 27. J Virol. 2022. PMID: 34705555 Free PMC article.
-
APOBEC3 proteins expressed in mammary epithelial cells are packaged into retroviruses and can restrict transmission of milk-borne virions.Cell Host Microbe. 2010 Dec 16;8(6):534-43. doi: 10.1016/j.chom.2010.11.003. Cell Host Microbe. 2010. PMID: 21147467 Free PMC article.
-
The receptor complex associated with human T-cell lymphotropic virus type 3 (HTLV-3) Env-mediated binding and entry is distinct from, but overlaps with, the receptor complexes of HTLV-1 and HTLV-2.J Virol. 2009 May;83(10):5244-55. doi: 10.1128/JVI.02285-08. Epub 2009 Mar 11. J Virol. 2009. PMID: 19279090 Free PMC article.
-
Genome-wide inference of protein interaction sites: lessons from the yeast high-quality negative protein-protein interaction dataset.Nucleic Acids Res. 2008 Apr;36(6):2002-11. doi: 10.1093/nar/gkn016. Epub 2008 Feb 14. Nucleic Acids Res. 2008. PMID: 18281313 Free PMC article.
References
-
- Bernard, K. A., W. B. Klimstra, and R. E. Johnston. 2000. Mutations in the E2 glycoprotein of Venezuelan equine encephalitis virus confer heparan sulfate interaction, low morbidity, and rapid clearance from blood of mice. Virology 276:93-103. - PubMed
-
- Bernstein, H. J. 2000. Recent changes to RasMol, recombining the variants. Trends Biochem. Sci. 25:453-455. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

