A novel human Ada2 homologue functions with Gcn5 or Brg1 to coactivate transcription
- PMID: 12972612
- PMCID: PMC193946
- DOI: 10.1128/MCB.23.19.6944-6957.2003
A novel human Ada2 homologue functions with Gcn5 or Brg1 to coactivate transcription
Abstract
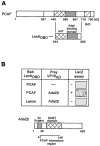
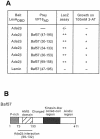
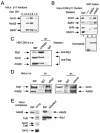
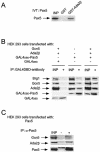
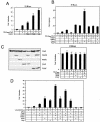
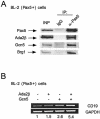
In yeast, the transcriptional adaptor yeast Ada2 (yAda2) is a part of the multicomponent SAGA complex, which possesses histone acetyltransferase activity through action of the yGcn5 catalytic enzyme. yAda2, among several SAGA proteins, serves to recruit SAGA to genes via interactions with promoter-bound transcription factors. Here we report identification of a new human Ada2 homologue, hAda2beta. Ada2beta differs both biochemically and functionally from the previously characterized hAda2alpha, which is a stable component of the human PCAF (human Gcn5 homologue) acetylase complex. Ada2beta, relative to Ada2alpha, interacted selectively, although not stably, with the Gcn5-containing histone acetylation complex TFTC/STAGA. In addition, Ada2beta interacted with Baf57 (a component of the human Swi/Snf complex) in a yeast two-hybrid screen and associated with human Swi/Snf in vitro. In functional assays, hAda2beta (but not Ada2alpha), working in concert with Gcn5 (but not PCAF) or Brg1 (the catalytic component of hSwi/Snf complex), increased transcription via the B-cell-specific transcription factor Pax5/BSAP. These findings support the view that Gcn5 and PCAF have distinct roles in vivo and suggest a new mechanism of coactivator function, in which a single adaptor protein (Ada2beta) can coordinate targeting of both histone acetylation and chromatin remodeling activities.
Figures


References
-
- Aasland, R., A. F. Stewart, and T. Gibson. 1996. The SANT domain: a putative DNA-binding domain in the SWI-SNF and ADA complexes, the transcriptional co-repressor N-CoR and TFIIIB. Trends Biochem. Sci. 21:87-88. - PubMed
-
- Barlev, N. A., R. Candau, L. Wang, P. Darpino, N. Silverman, and S. L. Berger. 1995. Characterization of physical interactions of the putative transcriptional adaptor, ADA2, with acidic activation domains and TATA-binding protein. J. Biol. Chem. 270:19337-19344. - PubMed
-
- Barlev, N. A., L. Liu, N. H. Chehab, K. Mansfield, K. G. Harris, T. D. Halazonetis, and S. L. Berger. 2001. Acetylation of p53 activates transcription through recruitment of coactivators/histone acetyltransferases. Mol. Cell 8:1243-1254. - PubMed
-
- Bell, B., E. Scheer, and L. Tora. 2001. Identification of hTAF(II)80 delta links apoptotic signaling pathways to transcription factor TFIID function. Mol. Cell 8:591-600. - PubMed
-
- Bell, B., and L. Tora. 1999. Regulation of gene expression by multiple forms of TFIID and other novel TAFII-containing complexes. Exp. Cell Res. 246:11-19. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
